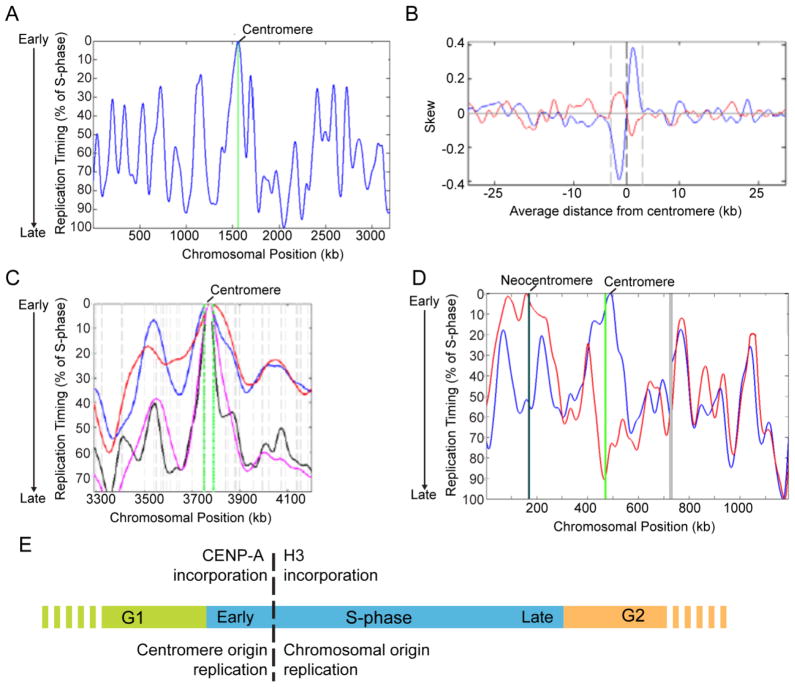
Figure 2. Efficient DNA replication at fungal centromeres.
(A) Centromeres are the earliest and most efficient origins on each C. albicans chromosome. The replication timing profile for chromosome in wild-type cells is indicated in blue. The position of CEN1 is indicated with a green line. (B) Centromere DNA in C. albicans and related species contains GC-skew, a replication-dependent strand-bias sequence pattern characteristic of constitutively active replication origins. Mean GC-skew (blue) and AT-skew (red) skew at the C. albicans centromere regions from all chromosomes are shown. (C) In S. pombe, CEN1 replicates earliest of all loci on chromosome 1. The centromere is indicated in green, while black, magenta, blue and red lines indicate data from different S. pombe replication timing microarray experiments (Feng et al., 2006, Heichinger et al., 2006, Mickle et al., 2007). (D) Neocentromere loci become the earliest, most efficient origins following kinetochore assembly, indicating that the replication pattern is determined by the presence of a function kinetochore. The native centromere (green line) is the earliest replicating region in wild-type cells (blue line), while the neocentromere locus approximately 170kb from the left telomere (black line) is the earliest replicating region in cells with this homozygous neocentromere (red line). Gray lines indicate a gap in data from the multiple repeat sequences in C. albicans on the right arm of chromosome 5. (E) Model for coordination of replication timing and maintenance of centromere specification. Adapted from (Koren et al., 2010).