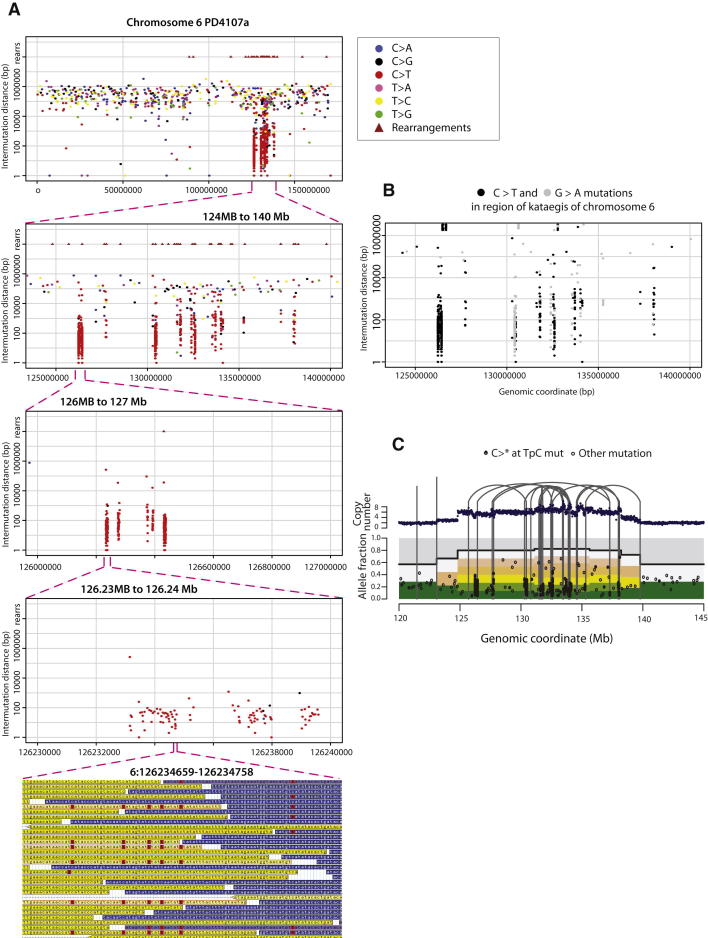
Figure 4.
Rainfall Plot for Chromosome 6 of PD4107a
(A) The x axis shows the genomic coordinates of the mutations. Rearrangements are presented as brown triangles (rearrs is an abbreviation for rearrangements). The region of kataegis is highlighted at increasing resolution to demonstrate microclusters within the macrocluster. The processive nature of C>T mutations at TpC context occurring in cis is seen in the lowest panel (G-browse image).
(B) Alternating processivity of kataegis in PD4107a. Long regions of C>T mutations are interspersed with regions of G>A mutations.
(C) Kataegis occurs with a variety of rearrangement architectures. Thick top line shows the copy number segments for the region of chromosome 6 of PD4107a. Point mutations are shown in lower panel as black points. x axis reflecting genomic position and y axis represents variant allele fraction. The proportions of reads derived from contaminating normal cells are depicted in gray and the fraction coming from each of the copies of that segment in the tumor cells are depicted by the multiple bars from green to yellow to pink to white. Early mutations will be found relatively higher up these bars, whereas late ones will be seen down the bottom of the variant allele fraction. Grey vertical lines represent rearrangements. Interconnecting lines indicate intrachromosomal rearrangements. On a macroscopic scale, this demonstrates how kataegis can be associated with chromothripsis (within region 130–135 Mb) as well as other rearrangement architectures.