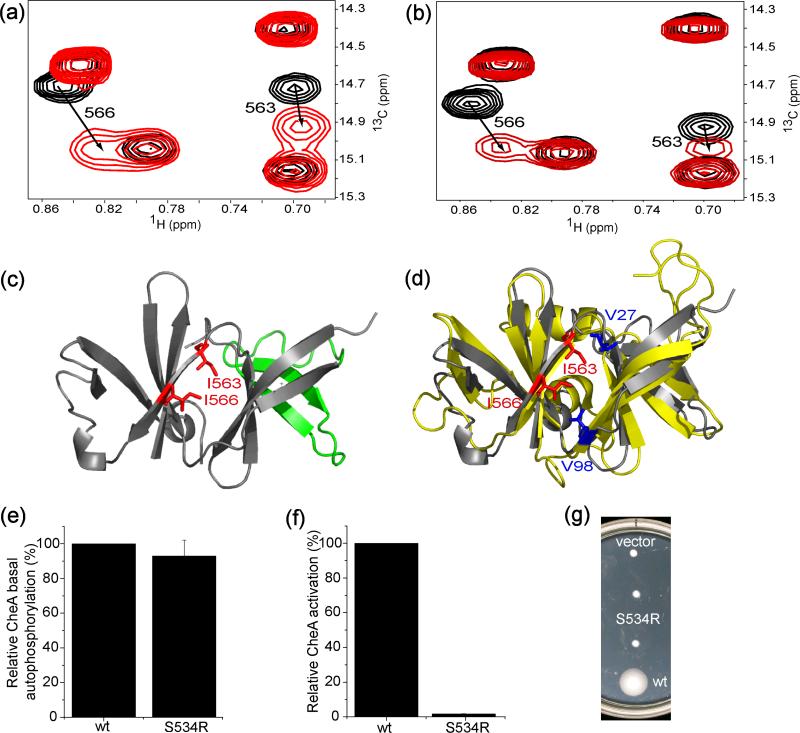
Fig. 2.
NMR and mutational analysis of the receptor binding site of CheA. (a) Methyl TROSY section of 13C-methyl labeled CheAΔ354 (150 μM) in the presence (red, at 1:4 ratio of CheAΔ354:TM001490-206) and absence (black) of TM001490-206. Residues with significant chemical shift changes are numbered with the arrows indicating the direction of the peak movement. (b) Same section of the spectra of methyl-labeled CheAΔ354 (150 μM) in complex with unlabeled CheW before (black) and after (red, 1:4 ratio of CheAΔ354-CheW: TM001490-206) the addition of TM001490-206. (c) Residues (red) showing significant chemical shift changes are mapped onto the structure of the P5 domain of CheA (PDB ID: 1B3Q). The region involved in CheW binding is colored in green13. (d) Structural alignment of CheW (yellow, PDB ID: 1K0S) and the P5 domain of CheA (grey). The two residues (Val27 and Val98) that show the largest shifts in the CheW-receptor interaction14 are highlighted in blue, and Ile563 and Ile566 that shift upon receptor binding are in red. (e) ATPase assays (s.d., n=3) for the basal autophosphorylation activities, (f) in vitro CheA activation assays (s.d., n=3) for the activation abilities, and (g) in vivo swarm assays for the chemotactic abilities of the wild-type E. coli CheA and S534R mutant (duplicate) at the receptor binding site of the P5 domain determined by NMR.