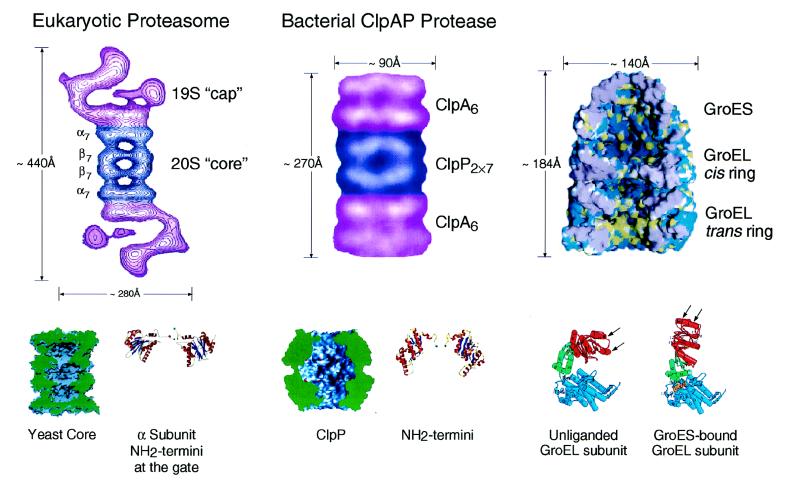
Figure 2.
Architecture of the eukaryotic proteasome and bacterial ClpAP chaperone–protease complexes and of the bacterial GroEL-GroES chaperonin pair. Side views from electron microscopy of the eukaryotic 26S proteasome (Left) and bacterial ClpAP (Center) showing the respective chaperone assemblies associated with the respective proteolytic cylinders (taken from ref. 11). The stoichiometries of the constitutent oligomeric rings are designated by subscripts; note that the eukaryotic proteasome is composed of seven distinct α subunits and seven distinct β subunits arranged 2-fold symmetrically to compose the four rings. Shown below are space-filling cutaway images of the proteolytic cylinders, derived from the crystal structures of Wang et al. (31) and Groll et al. (32), with active sites shown as red dots, as well as ribbon diagrams of their entryways, also taken from ref. 11. A space-filling view of the GroEL-GroES-ADP7 asymmetric chaperonin complex is shown (Upper Right), taken from Xu et al. (3), illustrating the differences between GroEL rings in the polypeptide-accepting and folding-active states. The open trans ring of the asymmetric complex exposes hydrophobic residues (shown in yellow) that can capture a non-native polypeptide. Subsequent GroES/ATP binding to the ring with polypeptide replaces this surface with a hydrophilic one (shown in blue), enlarges the cavity 2-fold in volume, and encapsulates the space in which a polypeptide, released from the hydrophobic binding sites, pursues folding in solitary confinement. Below, the rigid body movements of apical (red) and intermediate (green) domains of GroEL that occur on GroES binding are shown, taken from Xu et al. (3). The apical peptide binding surfaces of helices H and I (arrows), as well as an underlying segment, are removed from facing the central cavity to a position rotated upward 60° and twisted 90° clockwise (see text and ref. 3 for details).