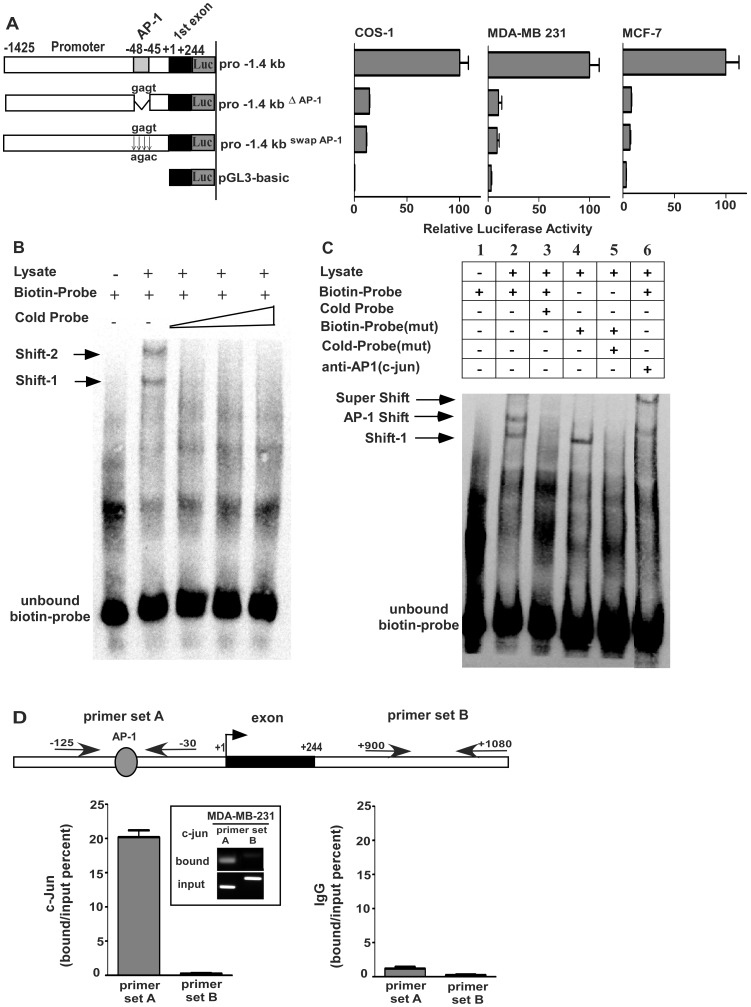
Figure 5. Requirement of AP-1 binding element in the KIAA1199 promoter.
A) A schematic diagram of mutations at the AP-1 binding site: A site-directed mutagenesis was carried out to generate either a deletion mutant by removing the AP-1 consensus sequence (GAGT) or a substitute mutation within the pro-1.4 promoter construct. The relative promoter activities of the mutations (ratio of firefly luciferase over Renilla luciferase) were then compared to the activity of the wild type pro-1.4 promoter construct (defined as arbitrary value of 100) in COS-1, MDA-MB-231, and MCF-7 cells. Error bars indicate mean +/− S.E. B) Binding of nuclear proteins to the AP-1 site in the KIAA1199 promoter: EMSA was carried out using a biotinylated double-stranded oligonucleotide (50 bp) containing the AP-1 binding site and nuclear extracts from MDA-MB-231 cells. Where indicated, binding was competed with 50–200 fold excess amounts of unlabeled probe. DNA-protein complexes formed are indicated as Shift-1 and Shift-2. C) Determination of specific binding between AP1 and the KIAA1199 promoter: EMSA was performed using a biotinylated double-stranded oligonucleotide (50 bp) containing the AP-1 binding site and nuclear extracts from MDA-MB-231 cells. Where indicated, biotinylated probe along with anti-C-Jun antibody or biotinylated probe containing mutated site of AP-1 consensus sequence were incubated with nuclear extracts from MDA-MB-231 cells. D) ChIP assay for analysis of association between endogenous AP-1 and the KIAA199 promoter sequence: A strong relation between AP-1 and DNA sequence was shown by 20% bound/input ratio as compared to the unrelated intron region. Normal rabbit IgG was used as a negative control. Results were calculated according to the bound/input ratio.