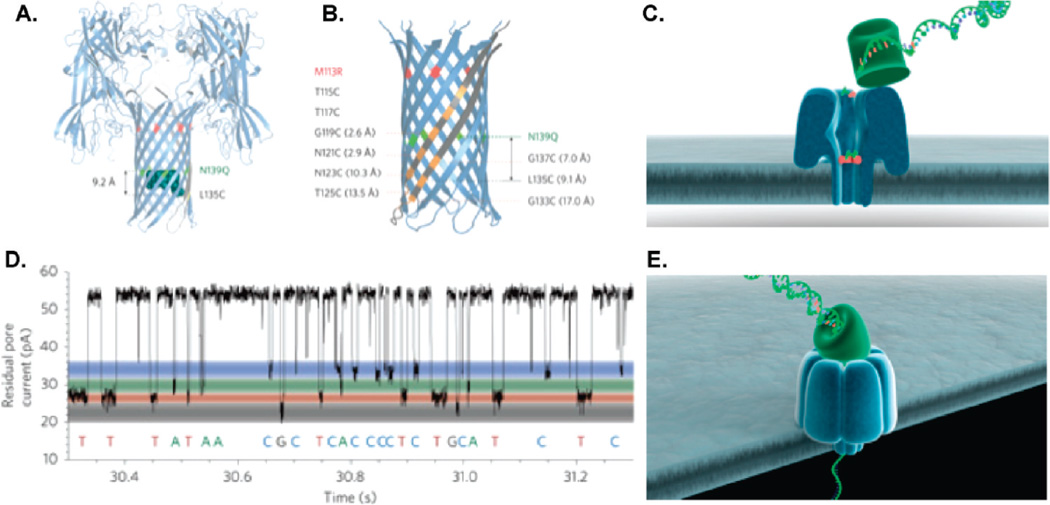
Figure 6.
Biological nanopore scheme employed by Oxford Nanopore. (A) Schematic of αHL protein nanopore mutant depicting the positions of the cyclodextrin (at residue 135) and glutamines (at residue 139). (B) A detailed view of the β barrel of the mutant nanopore shows the locations of the arginines (at residue 113) and the cysteines. (C) Exonuclease sequencing: A processive enzyme is attached to the top of the nanopore to cleave single nucleotides from the target DNA strand and pass them through the nanopore. (D) A residual current-vs-time signal trace from an αHL protein nanopore that shows a clear discrimination between single bases (dGMP, dTMP, dAMP, and dCMP). (E) Strand sequencing: ssDNA is threaded through a protein nanopore and individual bases are identified, as the strand remains intact. Panels A, B, and D reprinted with permission from ref 91. Copyright 2009 Nature Publishing Group. Panels C and E reprinted with permission from Oxford Nanopore Technologies (Zoe McDougall).