Figure 2.
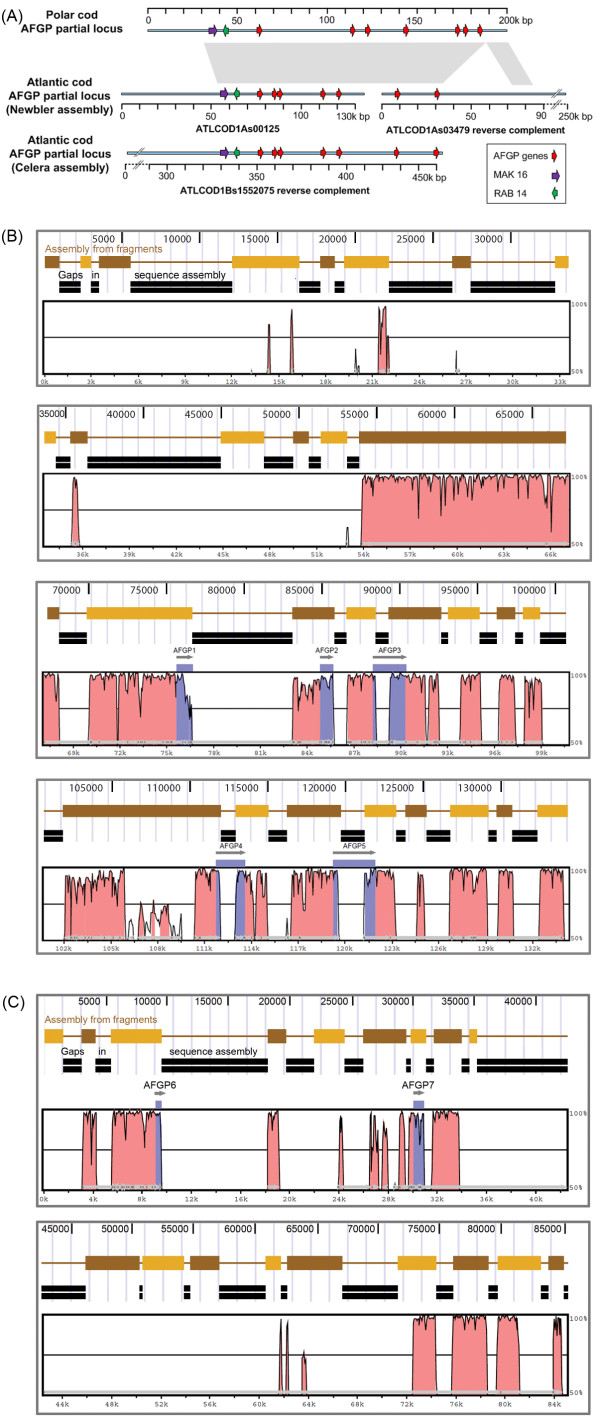
Alignment of AFGP-containing scaffolds in Atlantic cod and partial AFGP genomic locus from polar cod. (A) Schematic alignment map—Grey shaded areas indicate regions of high nucleotide identities between polar cod and Atlantic cod. AFGP partial locus of Atlantic cod is represented by two sequence scaffolds (ATLCOD1As00125 and ATLCOD1As03479) in the Newbler assembly (ATLCOD1A), and one sequence scaffold (ATLCOD1Bs1552075) in the Celera assembly (ATLCOD1B). AFGP genes of polar cod and fragmented coding sequences in the cod scaffolds are depicted as red arrows pointing towards the 3′ end. Hypothetical protein-coding genes MAK16-like and RAB14-like are denoted as purple and green arrows, respectively. (B, C) Nucleotide alignments determined using VISTA—The two Atlantic cod sequence scaffolds ATLCOD1As00125 (B) and ATLCOD1As03479 (C) are depicted as connected brown and orange rectangular bars (assembled sequence fragments) and double black bars (gaps between the sequence fragments), generated with UCSC Genome browser. The framed histogram below the Atlantic cod sequence scaffold structures was VISTA generated plots of their alignment with the polar cod partial AFGP locus sequence. Purple areas in the histogram denote conserved AFGP sequences, and grey arrows denote the AFGP genes we annotated in the two Atlantic cod scaffolds. Pink areas denote other conserved regions. In (B) the full length of ATLCOD1As00125 (134,272 bp) was shown. In (C), the first 85 kbp of ATLCOD1As03479 (254,310 bp, reverse complement sequence) was shown; the remaining region of the scaffold did not align with polar AFGP genomic region and therefore not shown. The conserved sequence blocks between Atlantic cod and polar shared very high sequence identities ranging from 80% to 99%, indicative of shared microsynteny.