Figure 2.
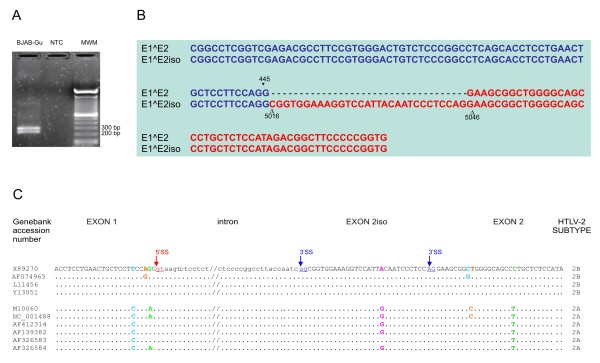
Identification of a novel HTLV-2 splice acceptor-site.A. Agarose gel electrophoresis of exon 1–2 and exon 1-2iso RT-PCR products. BJAB-Gu cDNA was amplified using the 6000-Gu forward primer (5’ GGCGGGCTCCTTCAACG 3’) located in the first exon and the 5183-Gu reverse primer (5’ CCATGGTGTTGGTGGTCT 3’) in the second exon. Amplification yielded the expected 275-bp band and a slightly longer product (lane 1). No bands were detected in the no template control (NTC, lane 2); lane 3 shows a 1-kbp molecular weight marker (MWM). Cell lines were cultured at 37°C, 5% CO2, in RPMI 1640 supplemented with 10% fetal bovine serum, 2 mM L-glutamine, penicillin (100 U/ml) and streptomycin (100 μg/ml). Cells were lysed in TRIzol® (Invitrogen) and total RNA was extracted and processed as detailed in Additional file 1: Table S1. B. Sequence alignment of cloned PCR products and identification of a novel splice acceptor site upstream of exon 2. Nucleotides in exon 1 are in blue and nucleotides in exon 2 are in red. Sequence analysis demonstrated that the 1-2iso junction is generated by splicing to a novel 3’splice-site (SS) located 30 nucleotides upstream of the exon 2 3'SS. Numbering of the sequence corresponds to that of the Gu isolate (GenBank: × 89270.1) in which + 1 is the first base of U3 in the 5’ LTR. ▼ indicates nucleotide position in exon 1; Δ indicates nucleotide position in exon 2 or 2iso. C. Sequence alignment of HTLV-2 isolates. DNA sequences containing exon 1 splice donor (5’SS) and exon 2iso or exon 2 splice-acceptor (3’SS) sites were compared between various HTLV-2 isolates retrieved from GenBank (accession numbers are indicated on the left, subtypes on the right). Exons and introns are indicated in upper and lower case, respectively. Underlined nucleotides indicate the 5’SS and 3’SS consensus sequences. The consensus sequence AGgtaagt at the exon 1-intron boundary varies among 2A and 2B subtypes, while the intronic gt 5’SS and ag 3’SS dinucleotides are invariable. The 3'SS defining exon 2iso is conserved among the different isolates examined.