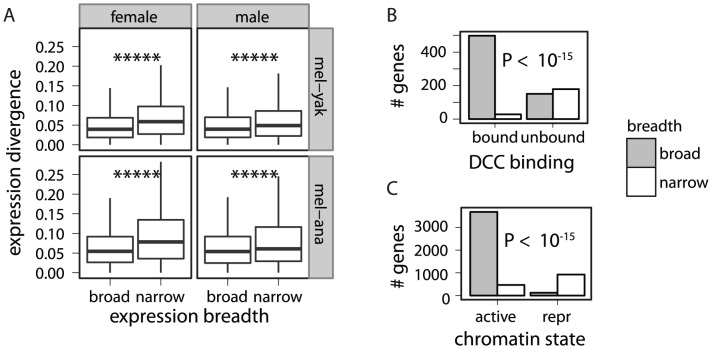
Figure 7. Association between expression breadth, DCC binding, chromatin state, and expression divergence.
Genes were classified based on their expression breadth, whether they are bound by the DCC, and whether they are in transcriptionally active or repressive (repr) chromatin (based on data from S2 cells). (A) Boxplots show the pairwise divergence in expression between 1∶1∶1 orthologs in the D. melanogaster (mel) and D. yakuba (yak) or D. ananassae (ana) genomes for broadly and narrowly expressed genes (see Figure 5). Mann-Whitney U tests were used to assess significant differences in expression divergence between broadly and narrowly expressed genes (***** 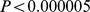 ). (B) X-linked broadly expressed (gray) and narrowly expressed (white) genes were divided into those that are bound and unbound by the DCC. (C) Broadly expressed (gray) and narrowly expressed (white) genes were divided into those that are in transcriptionally active and repressive chromatin. (B–C) Fisher's exact test was used to determine if there is a non-random distribution of genes in the four classes.
). (B) X-linked broadly expressed (gray) and narrowly expressed (white) genes were divided into those that are bound and unbound by the DCC. (C) Broadly expressed (gray) and narrowly expressed (white) genes were divided into those that are in transcriptionally active and repressive chromatin. (B–C) Fisher's exact test was used to determine if there is a non-random distribution of genes in the four classes.