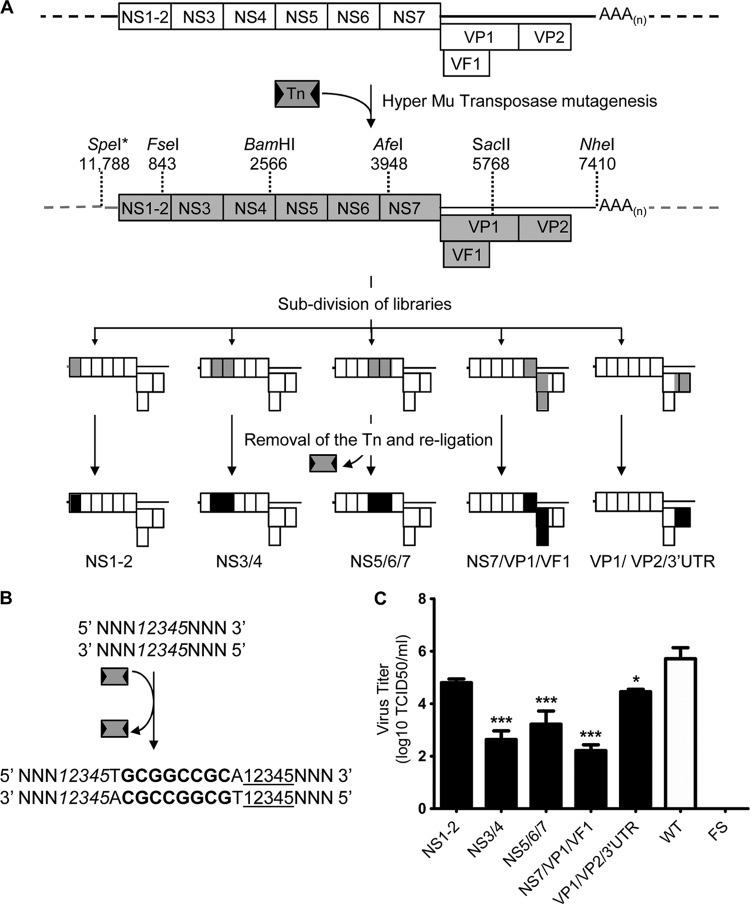
Fig 1.
(A) Strategy for Tn mutagenesis of the MNV-1 genome. A HyperMu transposase enzyme was used to insert one Tn per pT7:MNV 3′ RZ infectious clone to generate a Tn-mutagenized library. This was subcloned to generate five libraries containing Tns only in a well-defined region of the genome (gray shading).*, SpeI cleaves at a position within the plasmid vector; other numbers indicate genome positions. Removal of the Tns from each library leaves residual 15-nt insertions (black shading). Dashed lines, vector of the pT7:MNV 3′ RZ infectious clone. (B) Composition of the 15-nt insertion. Numbers in italics represent the 5-nt target site for the original transposition, 10 nt that originated from the transposon are shown and contain a NotI restriction site highlighted in bold, and the 5 underlined numbers represent target nucleotides duplicated during transposition. (C) Recovery of infectious virus from 15-nt insertion libraries. BSR-T7 cells were transfected with in vitro-transcribed, enzymatically capped RNA produced from each cDNA 15-nt insertion library. Virus was harvested at 48 h posttransfection, and the yield of recovered infectious virus was determined in RAW264.7 cells. The limit of detection is 50 TCID50/ml. Transfections were performed in triplicate; error bars represent the standard error of the mean. Significance was tested using one-way analysis of variance and Dunnett's multiple-comparison posttest to compare each insertion virus to the WT. FS, frameshift clone; ***, P < 0.001; *, P < 0.5.