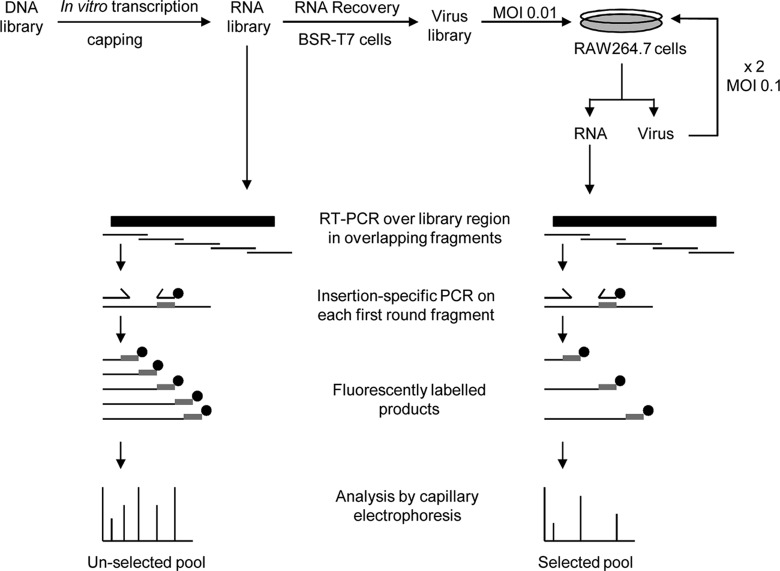
Fig 2.
Schematic diagram showing selection and detection of insertions in the 15-nt insertion library. RNA was produced from each 15-nt cDNA insertion library by in vitro transcription and enzymatic capping of the DNA library, and virus was recovered using the RNA-mediated reverse genetics system, in duplicate. For selection in tissue culture, the recovered virus was sequentially passed (in duplicate) three times in RAW264.7 cells at a low MOI. RNA was extracted after each passage and alongside the input RNA was used as a template for RT-PCR amplification of overlapping fragments covering the respective mutagenized region of each library. Each fragment was used as a template in a second round of PCR with a virus-specific primer and the insertion-specific NotI miniprimer, labeled with a fluorescent tag (black circles). Fluorescently labeled products were detected and analyzed by capillary electrophoresis.