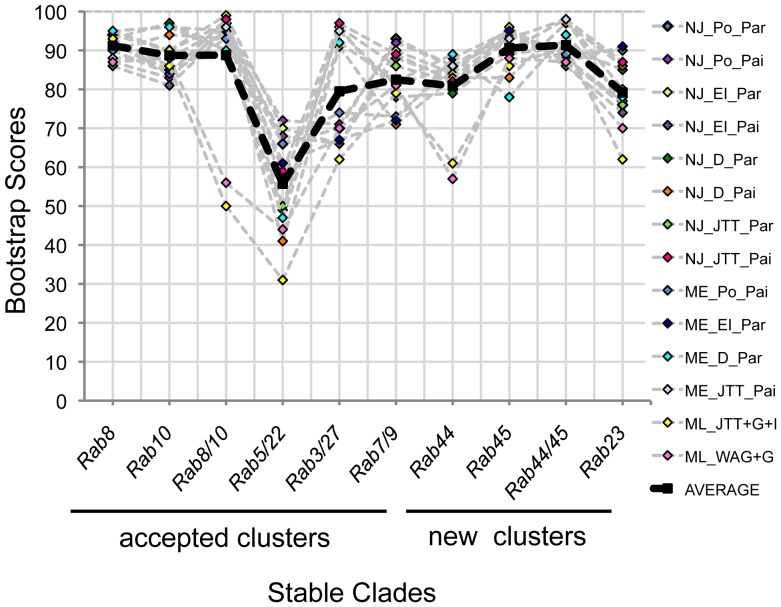
Figure 3. Bootstrap scores for specific terminal clades from 13 additional phylogenetic reconstructions.
Thirteen additional phylogenetic analyses were performed using a combination of statistical methods. Phylogenetic reconstruction methods include Neighbor Joining (NJ), maximum likelihood (ML) or minimum evolution (ME). Amino acid substitution methods include Poisson (Po), JTT, Dayhoff (D), Equal Input (EI) and WAG. Gap deletion treatments include partial (Par) or pairwise (Pai) and rates and patterns of evolution include gamma distributed (+G), invariant sites (+I) or uniform (all others). All phylogenetic reconstructions were performed in MEGA5. Specific orthologous and/or paralogous clades include both worm and human Rab proteins. New orthologous clusters include bootstrap scores supporting the Rab44/4R79.2, Rab45/C33D12.6 and Rab23/ZK669.5 pairs. The new paralogous cluster includes the bootstrap scores that support the Rab44,4R79.2,Rab45 and C33D12.6 terminal clade.