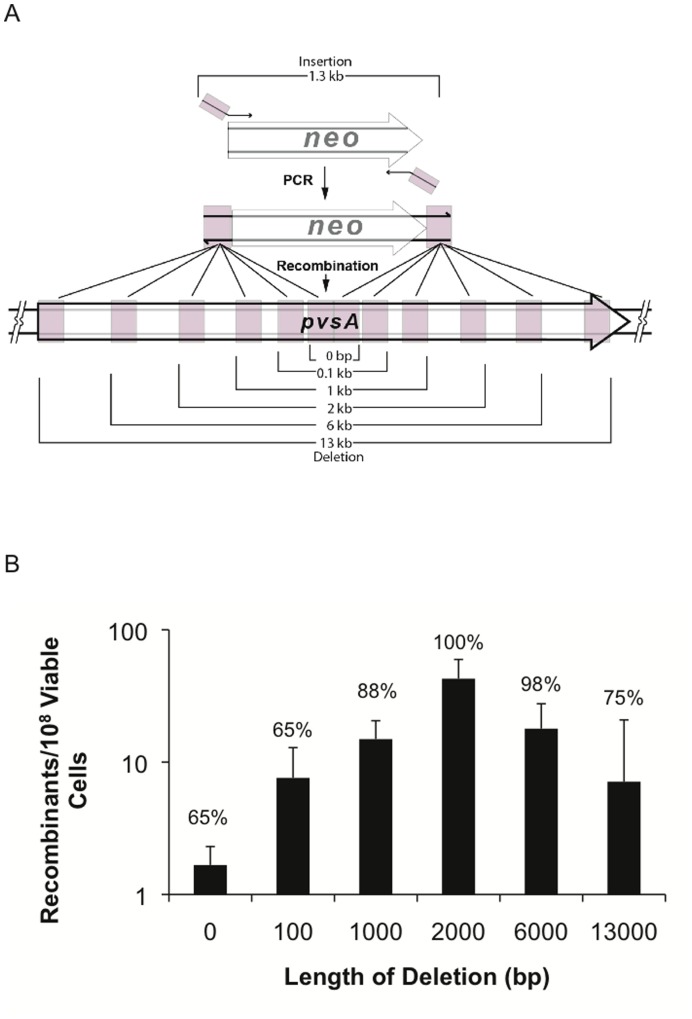
Figure 4. Deletion size affects RecTEPsy-mediated recombination efficiency.
(A) Recombination substrates were generated using PCR primers to amplify the neo gene with 80 nt homologies to the pvsA gene at the ends of the PCR product. The size of the deletion is determined by the distance between the pvsA flanking sequences as they exist on the P. syringae DC3000 genome. Homologous sequences are shown in pink. 500 ng of each substrate was used in electroporations. (B) Recombineering efficiency was determined for each substrate. Between 20 to 60 kanamycin resistant clones from each deletion length were analyzed using PCR to determine whether the neo gene had integrated and produced a deletion in the correct location. The percentage of kanamycin resistant clones with the correct deletion is indicated above each bar. The results are the average of four independent replicates, error bars indicate standard deviation.