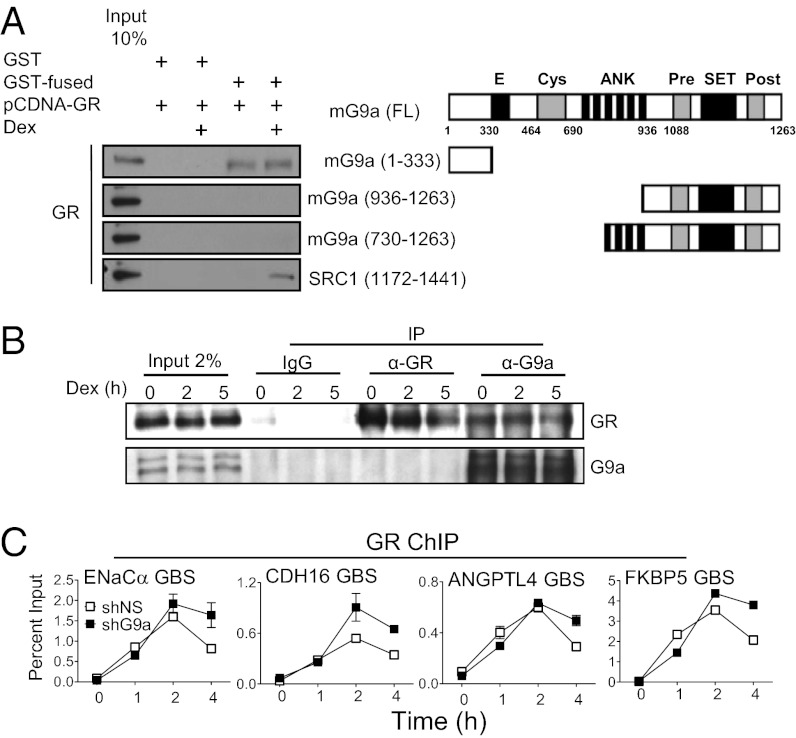
Fig. 3.
G9a interacts with GR in vitro and in vivo. (A) Right: Diagram of full-length mG9a (FL) and mG9a fragments used in this experiment showing amino acid sequence numbers and specific domains: Glu-rich (marked with “E”), Cys-rich ring finger-like (Cys), ankyrin repeat (ANK), methyltransferase (SET), and Cys-rich Pre-SET (Pre) and post-SET (Post). Left: Full-length human GR was synthesized in vitro, incubated with 1 µM dex or vehicle for 4 h, and incubated with GST or GST fused to mG9a fragments or to an SRC1a C-terminal fragment bound to glutathione-agarose beads as indicated. Bound human GR protein was detected by immunoblot with anti-GR antibody. A 10% input sample was loaded for comparison. (B) Coimmunoprecipitation of endogenous GR with G9a from lysates of A549 cells treated with 100 nM dex for the indicated times (in hours). Immunoprecipitation was performed with anti-G9a, anti-GR, or control IgG antibodies. Western blots were performed with anti-GR (Upper) and anti-G9a (Lower) antibodies. A 2% input sample was loaded for comparison. (C) Effect of G9a depletion on GR binding to target genes. ChIP was performed on A549 cells expressing shNS (open symbols) or shG9a (filled symbols) and treated with 100 nM dex for the indicated times. Immunoprecipitated DNA was analyzed by qPCR by using primers that amplify the GBS region of the indicated GR target genes. Results shown are mean ± SD of PCR reactions (n = 3) performed with DNA samples from a single experiment, and are representative of three independent experiments.