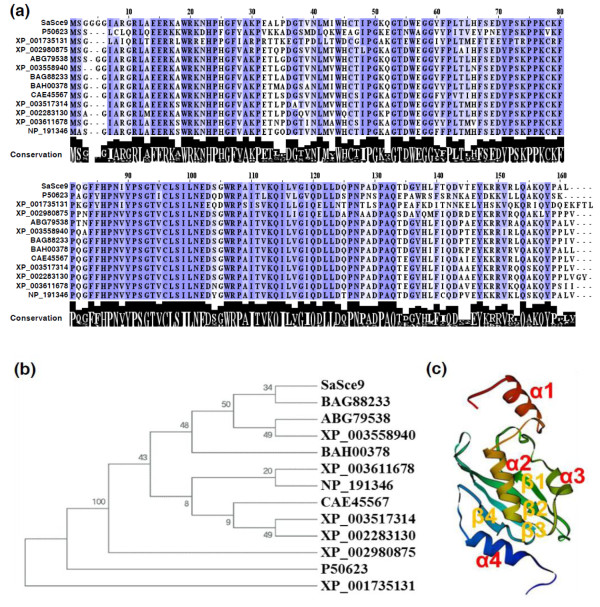
Figure 1.
Multiple sequence alignments, phylogenetic analysis, and predicted tertiary structure of SaSce9 protein. (a) Multiple sequence alignment of SaSce9 protein with SCE proteins from various organisms. Conservation of amino acid residues are shown by black bars below the alignments. Accession numbers of sequences for SCE proteins are: P50623 (Saccharomyces cerevisiae), XP_001735131 (Entamoeba histolytica), XP_002980875 (Selaginella moellendorffii), ABG79538 (Triticum durum), XP_003558940 (Brachypodium distachyon), BAG88233 {Oryza sativa (Os03g0123100)}, BAH00378 {Oryza sativa (Os10g0536000)}, CAE45567 (Nicotiana benthamiana), XP_003517314 (Glycine max), XP_002283130 (Vitis vinifera), XP_003611678 (Medicago truncatula), NP_191346 (Arabidopsis thaliana); (b) Phylogenetic tree of SaSce9. The amino acid sequences were subjected to Bootstrap test of phylogeny by the MEGA 4.0 program, using neighbour-joining method with 1000 replicates. Numbers on the Figure are bootstrap values; (c) Model of predicted tertiary structure was performed using SWISS-MODEL based on crystallographic data deposited on the Swissprot.