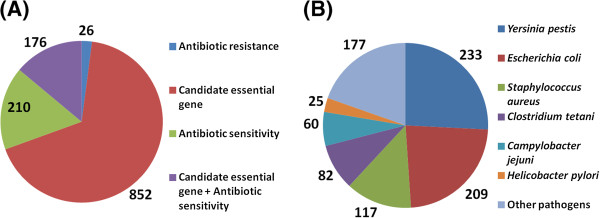
Figure 3.
Missed genes that can be associated with COMBREX phenotype data: (A) Phenotype data distribution. Some of the missed genes can be associated with phenotype data using the novel COMBREX resource. A gene is associated with a specific phenotype if it has significant sequence similarity (see Methods) to a gene in COMBREX with the phenotype or if it is assigned to a cluster containing a gene with the phenotype. (B) Potential drug target genes (see text for details) with their distribution in pathogenic organisms (identified only to the species level). The largest portion of these genes belongs to the Enterobacteriaceae family of pathogens (Yersinia, E. coli, etc.) as seen in the figure. This phenotype information is pooled together using ComBlast. The full list of these genes is available online [13].