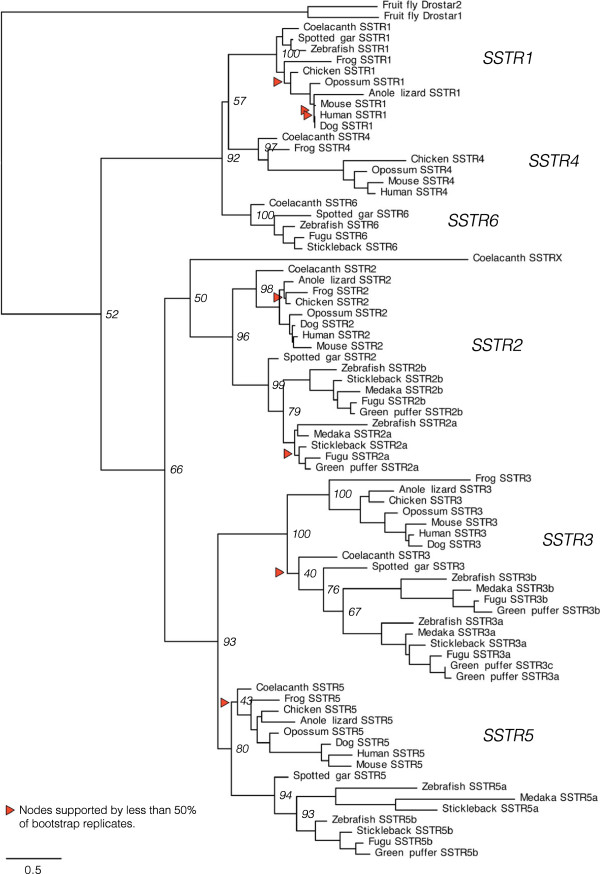
Figure 1.
Phylogenetic maximum likelihood tree of the somatostatin receptor gene family. The topology is supported by a non-parametric bootstrap test with 100 replicates as well as an SH-like approximate likelihood ratio test (aLRT). The tree is rooted with the human kisspeptin receptor 1 sequence (not shown). Branch support (bootstrap replicates) for deep divergences is shown at the nodes. All branch support values are shown in Figure S1 (bootstrap replicates) and Figure S2 (aLRT) (see Additional file 2). The phylogenetic tree shows six well-supported subtype clusters, with the somatostatin receptor subtypes SSTR2, -3 and -5 forming one ancestral branch and the SSTR1, -4 and -6 receptor subtypes forming one ancestral branch. This phylogenetic analysis supports the emergence of all six subtypes early in vertebrate evolution, with the subsequent loss of SSTR4 in ray-finned fishes, before the divergence of the spotted gar and teleost lineages, and of SSTR6 in the tetrapod lineage. All six subtypes could be identified in the coelacanth genome. A seventh SSTR2-like sequence, called SSTRX in the tree, could also be identified on the same genomic scaffold in the coelacanth genome (see Additional file 1, Supplemental note 1). There are well-supported teleost-specific duplicate branches of SSTR2, -3 and -5, although all could not be identified in all teleost genomes. These duplicates have been named a and b based on the phylogenetic analysis. There is a third SSTR3 sequence in the green puffer, called SSTR3c in the tree.