Figure 3.
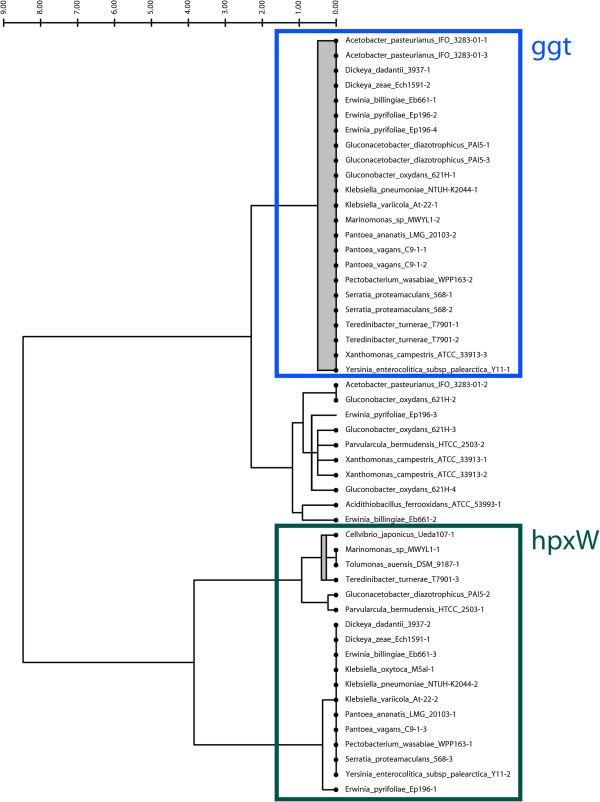
hpxW and ggt context tree. Clusters of all homologous gene clusters in 22 alpha and gamma proteobacterial species were constructed using BLAST [33] and tribe-MCL [34]. All ggt and hpxW genes naturally grouped into the same homology cluster. Using JContextExplorer, we defined a context set, which we named “D75”, that placed all genes on the same strand within 75 nucleotides of each other into common gene groupings. We constructed a context tree of the ggt/hpxW homology cluster using the “Common Genes – Dice” dissimilarity metric and “Joint Between-Within” linkage function (above). The data segmented into two branches, one of which corresponded to the previously described hpxW context (green box), and the other into a combination of the ggt context (blue box) and an undetermined third group. Manual inspection of individual contexts in the third group might reveal that some members of this group belong with the ggt group, and some with the hpxW group. Additionally, some members in this third unknown group could represent “transitional” cases between the hpxW and ggt gene (a gene that performs the functions of both hpxW and ggt, for example). JContextExplorer’s context viewer tool proved helpful in manually interrogating this third group.