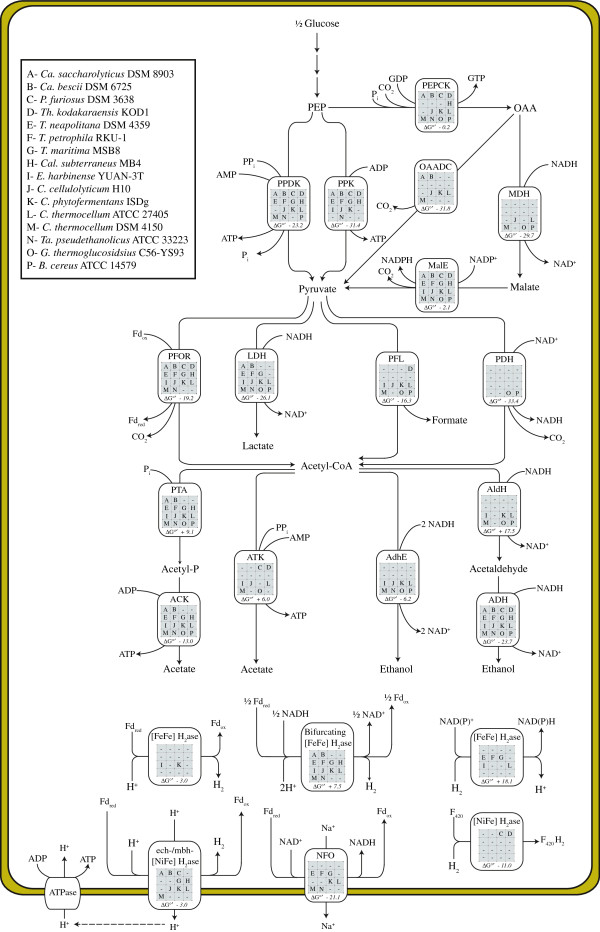
Figure 1.
Comparison of putative gene products involved in pyruvate metabolism and end-product synthesis among select hydrogen and ethanol-producing species. Presence of putative gene products are indicated in matrix with respective letters corresponding to selected organism (see legend). Numbers indicate standard free energies of reaction (△G°’) corresponding to a particular enzyme. Abbreviations: PEPCK, phosphoenolpyruvate carboxykinase; OAADC, oxaloacetate decarboxylase; MDH, malate dehydrogenase; MalE, malic enzyme; PPK, pyruvate kinase; PPDK, pyruvate phosphate dikinase; LDH, lactate dehydrogenase; PFL, pyruvate formate lyase; PFOR, pyruvate:ferredoxin oxidoreductase; PDH, pyruvate dehydrogenase; ADH, alcohol dehydrogenase; ALDH, acetaldehyde dehydrogenase; AdhE, bifinctional acetaldehyde/alcohol dehydrogenase; ACK, acetate kinase; PTA, phosphotransacetylase; NFO, NADH:Fd oxidoreductase.