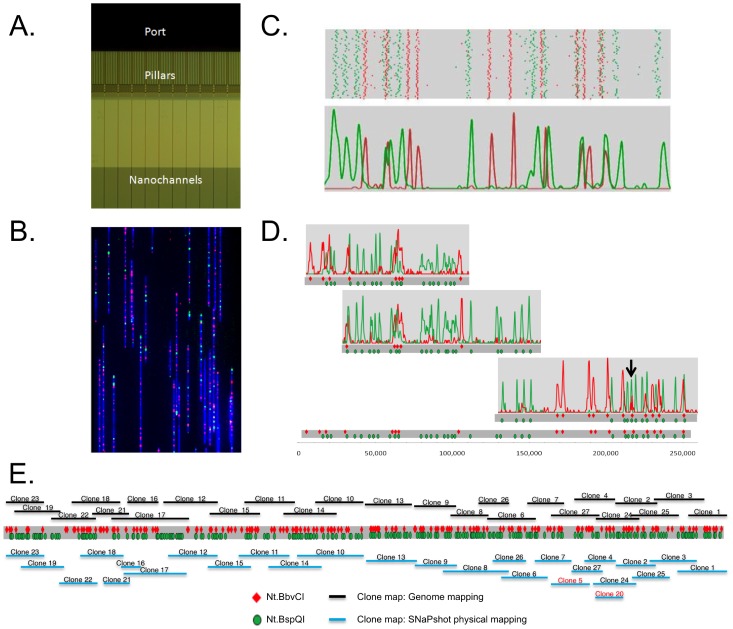
Figure 1. Two-color genome mapping with two enzymes.
A. The DNA backbone is stained with YOYO-1 and loaded into the port of a nanochannel array chip. The DNA molecules are introduced into the region with pillars and micron-scale relaxation channels by an electric field where they unwind and linearize. Finally, they are moved into the 45 nm nanochannels, where they stretch uniformly to 85% of the length of perfectly linear B-DNA. B. Linearized BAC DNA molecules in nanochannels. The DNA molecule is stained with YOYO-1, and Nt.BspQI and Nt.BbvCI nicks are labeled with green and red dyes, respectively. C. Molecule length and nick locations are extracted from the images by custom image-analysis software. By clustering individual molecules with high similarity of green label patterns, distinct patterns are extracted (top panel). The locations of the red labels are then overlaid on the green label patterns (middle pattern). A histogram plot of the above clusters is shown in the bottom panel. The peaks represent the location of each sequence motif (GCTCTTC and CCTCAGC) along the linearized DNA molecules. D. Consensus maps for individual BAC clones are shown. Consensus maps are combined by using overlapping patterns, and the final genome map is shown at the bottom. E. The clone map from genome mapping is shown at the top and the full genome map as a grey bar with Nt.BspQI and Nt.BbvCI motif locations in green and red. Below the genome map, in blue, is the physical map from SNaPshot fingerprinting. The total length of the genome map is 2.1 Mb.