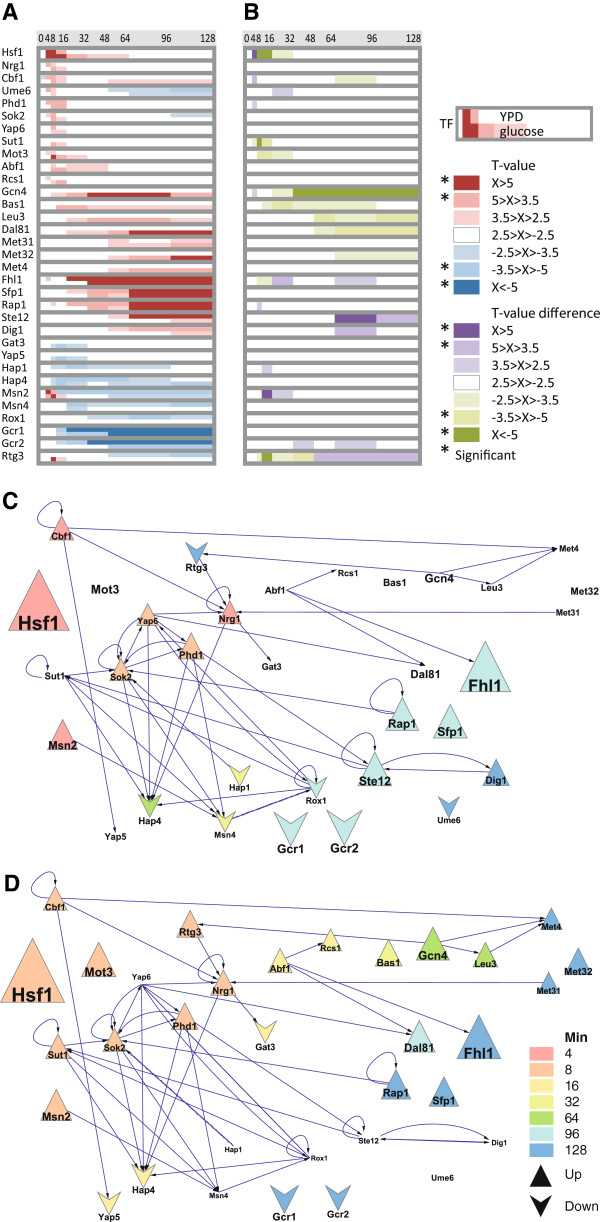
Figure 4.
Differences in expression regulation between YPD and glucose induced germination include the activity of TFs for amino acid biosynthesis. (A) T-profiler prediction of TFs involved in the germination on YPD and glucose based on ChIP-chip results [20]. Colours indicate the relative TF activity (T-values) of YPD (upper row) and glucose (lower row) induced germination for each TF according to panel. (B) Differences in T-values for YPD and glucose germination highlight the differential induction of the amino acid biosynthetic subprogram (in glucose only) and timing of mating (faster in YPD). Differences of ±2.5 or more are indicated in purple (stronger in YPD) and green (stronger in glucose), and scaled according to the colour panel. Note that this colour indicates largest deviation from zero, not the highest value. White (and weak colours) indicate no significant difference. (C, D) The germination program is orchestrated by an interconnected network of TFs that are activated sequentially. TF node size indicates activity (T-profiler score), node colour indicates time of peak activity (see legend) and node shape indicates directionality (up or down). Edges show cross regulation within the TF network, i.e. reported TF binding to promoter of (other) TF [21]. Note that the timing is different in YPD (C) and glucose (D), and that different TFs are activated (missing nodes show no significant activity under that condition).