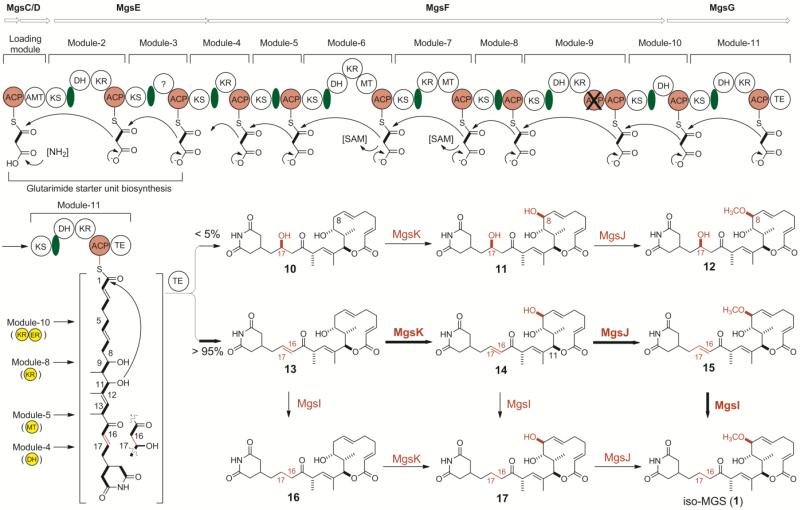
Figure 2.
Proposed biosynthetic pathway for iso-MGS (1) featuring the iso-MGS AT-less type I PKS that lacks a dehydratase (DH) domain in module-4, a MT domain in module-5, a KR domain in module-8, and a KR and an ER domain in module-10 according to the co-linear PKS model for the biosynthesis of the two nascent products, 10 and 13, and the tailoring enzymes MgsIJK with broad substrate specificity that convert 10 to 12 and 13 to 1. Biosynthesis of the glutarimide starter unit by the loading module, module-2 and module-3 has been previously proposed.10a Heavy arrows denote the preferred pathway by the tailoring enzymes with thin arrows accounting for metabolites resulted from substrate promiscuity. ACP, acyl carrier protein; AMT, amidotransferase; DH, dehydratase; ER, enoylreductase; KR, ketoreductase; KS, ketosynthase; MT, methyltransferase; TE, thioesterase. [NH2] depicts the amino donor for the AMT domain, [SAM] denotes S-adenosylmethionine as the methyl donor for the MT domain, the ACP domain with an overlaid X is nonfunctional, the domain with ? denotes an unknown function, green ovals depict AT-docking domains, and the yellow domains highlight the missing domains for the iso-MGS AT-less type I PKS predicted according to the co-linear model for the biosynthesis of 10 and 13.10a,11