Abstract
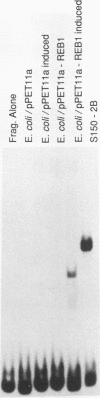
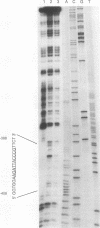
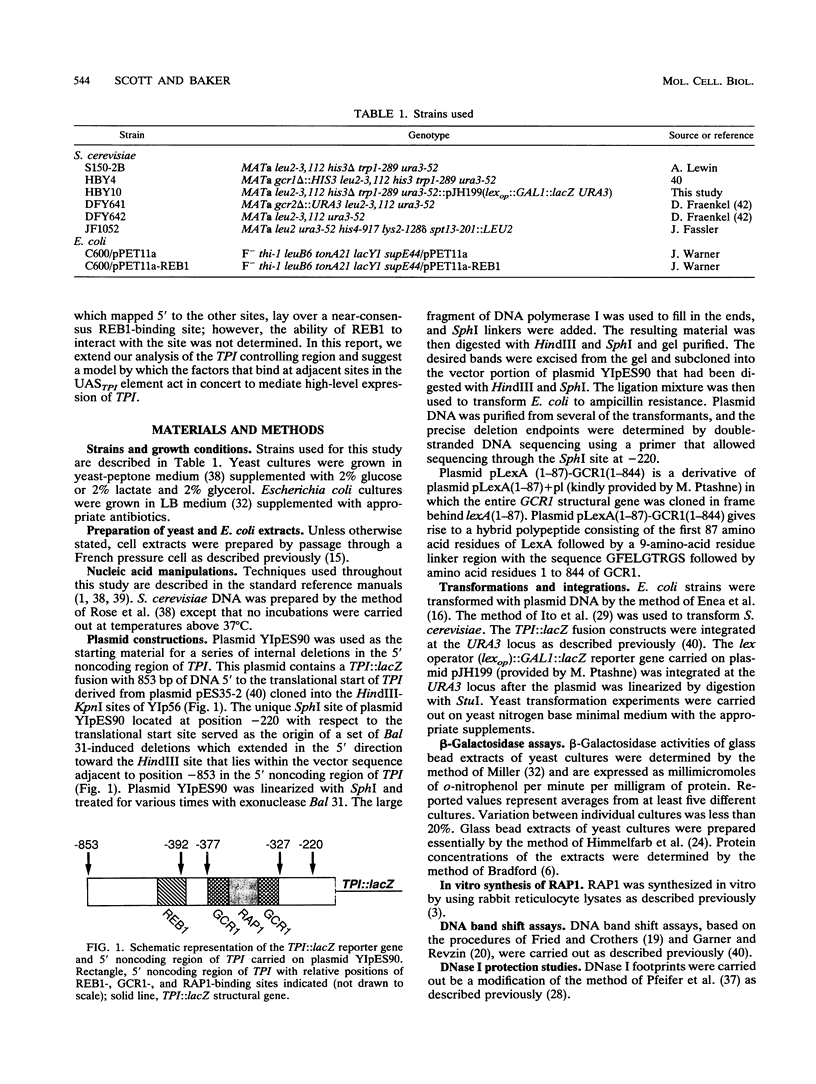
In Saccharomyces cerevisiae, the TPI gene product, triosephosphate isomerase, makes up about 2% of the soluble cellular protein. Using in vitro and in vivo footprinting techniques, we have identified four binding sites for three factors in the 5' noncoding region of TPI: a REB1-binding site located at positions -401 to -392, two GCR1-binding sites located at positions -381 to -366 and -341 to -326, and a RAP1-binding site located at positions -358 to -346. We tested the effects of mutations at each of these binding sites on the expression of a TPI::lacZ gene fusion which carried 853 bp of the TPI 5' noncoding region integrated at the URA3 locus. The REB1-binding site is dispensable when material 5' to it is deleted; however, if the sequence 5' to the REB1-binding site is from the TPI locus, expression is reduced fivefold when the site is mutated. Because REB1 blocks nucleosome formation, the most likely function of its binding site in the TPI controlling region is to prevent the formation of nucleosomes over the TPI upstream activation sequence. Mutations in the RAP1-binding site resulted in a 10-fold reduction in expression of the reporter gene. Mutating either GCR1-binding site alone had a modest effect on expression of the fusion. However, mutating both GCR1-binding sites resulted in a 68-fold reduction in the level of expression of the reporter gene. A LexA-GCR1 fusion protein containing the DNA-binding domain of LexA fused to the amino terminus of GCR1 was able to activate expression of a lex operator::GAL1::lacZ reporter gene 116-fold over background levels. From this experiment, we conclude that GCR1 is able to activate gene expression in the absence of REB1 or RAP1 bound at adjacent binding sites. On the basis of these results, we suggest that GCR1 binding is required for activation of TPI and other GCR1-dependent genes and that the primary role of other factors which bind adjacent to GCR1-binding sites is to facilitate of modulate GCR1 binding in vivo.
Full text
PDF


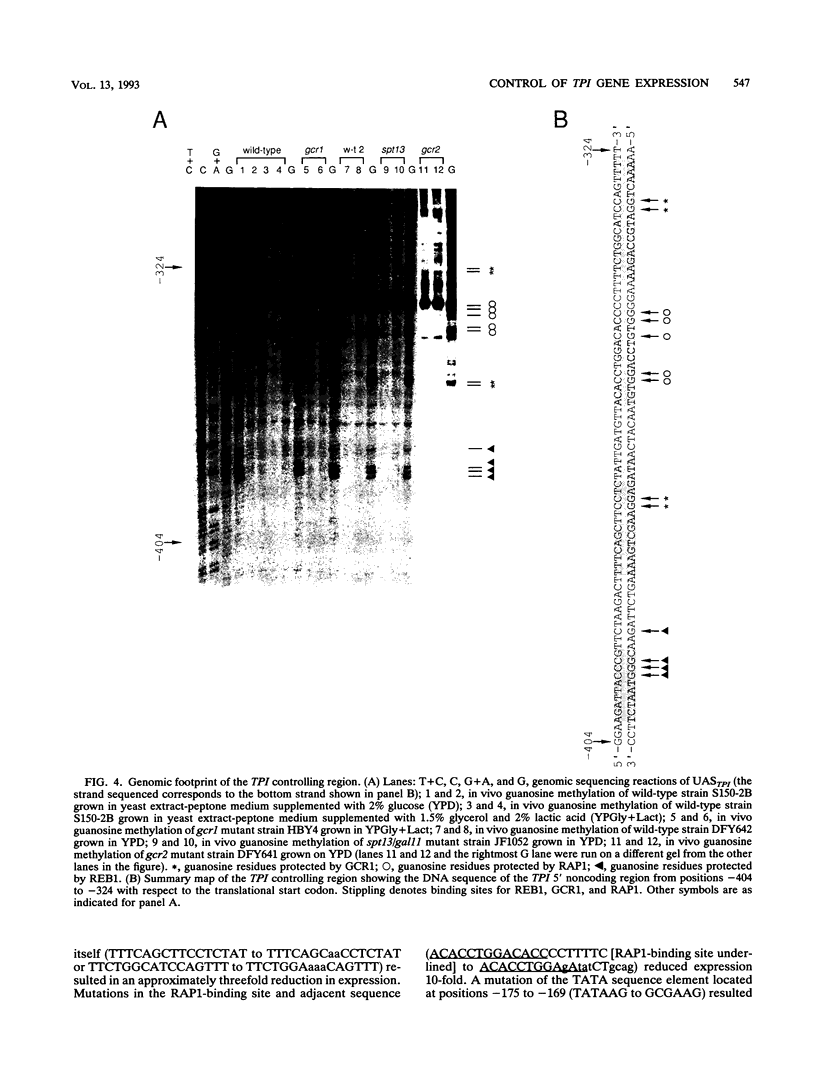



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baker H. V. GCR1 of Saccharomyces cerevisiae encodes a DNA binding protein whose binding is abolished by mutations in the CTTCC sequence motif. Proc Natl Acad Sci U S A. 1991 Nov 1;88(21):9443–9447. doi: 10.1073/pnas.88.21.9443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker H. V. Glycolytic gene expression in Saccharomyces cerevisiae: nucleotide sequence of GCR1, null mutants, and evidence for expression. Mol Cell Biol. 1986 Nov;6(11):3774–3784. doi: 10.1128/mcb.6.11.3774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bitter G. A., Chang K. K., Egan K. M. A multi-component upstream activation sequence of the Saccharomyces cerevisiae glyceraldehyde-3-phosphate dehydrogenase gene promoter. Mol Gen Genet. 1991 Dec;231(1):22–32. doi: 10.1007/BF00293817. [DOI] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1006/abio.1976.9999. [DOI] [PubMed] [Google Scholar]
- Brandl C. J., Struhl K. A nucleosome-positioning sequence is required for GCN4 to activate transcription in the absence of a TATA element. Mol Cell Biol. 1990 Aug;10(8):4256–4265. doi: 10.1128/mcb.10.8.4256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brent R., Ptashne M. A eukaryotic transcriptional activator bearing the DNA specificity of a prokaryotic repressor. Cell. 1985 Dec;43(3 Pt 2):729–736. doi: 10.1016/0092-8674(85)90246-6. [DOI] [PubMed] [Google Scholar]
- Brindle P. K., Holland J. P., Willett C. E., Innis M. A., Holland M. J. Multiple factors bind the upstream activation sites of the yeast enolase genes ENO1 and ENO2: ABFI protein, like repressor activator protein RAP1, binds cis-acting sequences which modulate repression or activation of transcription. Mol Cell Biol. 1990 Sep;10(9):4872–4885. doi: 10.1128/mcb.10.9.4872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butler G., Dawes I. W., McConnell D. J. TUF factor binds to the upstream region of the pyruvate decarboxylase structural gene (PDC1) of Saccharomyces cerevisiae. Mol Gen Genet. 1990 Sep;223(3):449–456. doi: 10.1007/BF00264453. [DOI] [PubMed] [Google Scholar]
- Chambers A., Stanway C., Tsang J. S., Henry Y., Kingsman A. J., Kingsman S. M. ARS binding factor 1 binds adjacent to RAP1 at the UASs of the yeast glycolytic genes PGK and PYK1. Nucleic Acids Res. 1990 Sep 25;18(18):5393–5399. doi: 10.1093/nar/18.18.5393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers A., Tsang J. S., Stanway C., Kingsman A. J., Kingsman S. M. Transcriptional control of the Saccharomyces cerevisiae PGK gene by RAP1. Mol Cell Biol. 1989 Dec;9(12):5516–5524. doi: 10.1128/mcb.9.12.5516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chasman D. I., Lue N. F., Buchman A. R., LaPointe J. W., Lorch Y., Kornberg R. D. A yeast protein that influences the chromatin structure of UASG and functions as a powerful auxiliary gene activator. Genes Dev. 1990 Apr;4(4):503–514. doi: 10.1101/gad.4.4.503. [DOI] [PubMed] [Google Scholar]
- Clifton D., Fraenkel D. G. The gcr (glycolysis regulation) mutation of Saccharomyces cerevisiae. J Biol Chem. 1981 Dec 25;256(24):13074–13078. [PubMed] [Google Scholar]
- Clifton D., Weinstock S. B., Fraenkel D. G. Glycolysis mutants in Saccharomyces cerevisiae. Genetics. 1978 Jan;88(1):1–11. doi: 10.1093/genetics/88.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Enea V., Vovis G. F., Zinder N. D. Genetic studies with heteroduplex DNA of bacteriophage fl. Asymmetric segregation, base correction and implications for the mechanism of genetic recombination. J Mol Biol. 1975 Aug 15;96(3):495–509. doi: 10.1016/0022-2836(75)90175-8. [DOI] [PubMed] [Google Scholar]
- Fedor M. J., Lue N. F., Kornberg R. D. Statistical positioning of nucleosomes by specific protein-binding to an upstream activating sequence in yeast. J Mol Biol. 1988 Nov 5;204(1):109–127. doi: 10.1016/0022-2836(88)90603-1. [DOI] [PubMed] [Google Scholar]
- Fried M., Crothers D. M. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 1981 Dec 11;9(23):6505–6525. doi: 10.1093/nar/9.23.6505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner M. M., Revzin A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 1981 Jul 10;9(13):3047–3060. doi: 10.1093/nar/9.13.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gill G., Ptashne M. Negative effect of the transcriptional activator GAL4. Nature. 1988 Aug 25;334(6184):721–724. doi: 10.1038/334721a0. [DOI] [PubMed] [Google Scholar]
- Golemis E. A., Brent R. Fused protein domains inhibit DNA binding by LexA. Mol Cell Biol. 1992 Jul;12(7):3006–3014. doi: 10.1128/mcb.12.7.3006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hess B., Boiteux A., Krüger J. Cooperation of glycolytic enzymes. Adv Enzyme Regul. 1969;7:149–167. doi: 10.1016/0065-2571(69)90016-8. [DOI] [PubMed] [Google Scholar]
- Himmelfarb H. J., Pearlberg J., Last D. H., Ptashne M. GAL11P: a yeast mutation that potentiates the effect of weak GAL4-derived activators. Cell. 1990 Dec 21;63(6):1299–1309. doi: 10.1016/0092-8674(90)90425-e. [DOI] [PubMed] [Google Scholar]
- Holland M. J., Holland J. P. Isolation and identification of yeast messenger ribonucleic acids coding for enolase, glyceraldehyde-3-phosphate dehydrogenase, and phosphoglycerate kinase. Biochemistry. 1978 Nov 14;17(23):4900–4907. doi: 10.1021/bi00616a007. [DOI] [PubMed] [Google Scholar]
- Holland M. J., Yokoi T., Holland J. P., Myambo K., Innis M. A. The GCR1 gene encodes a positive transcriptional regulator of the enolase and glyceraldehyde-3-phosphate dehydrogenase gene families in Saccharomyces cerevisiae. Mol Cell Biol. 1987 Feb;7(2):813–820. doi: 10.1128/mcb.7.2.813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huie M. A., Scott E. W., Drazinic C. M., Lopez M. C., Hornstra I. K., Yang T. P., Baker H. V. Characterization of the DNA-binding activity of GCR1: in vivo evidence for two GCR1-binding sites in the upstream activating sequence of TPI of Saccharomyces cerevisiae. Mol Cell Biol. 1992 Jun;12(6):2690–2700. doi: 10.1128/mcb.12.6.2690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Machida M., Jigami Y., Tanaka H. Purification and characterization of a nuclear factor which binds specifically to the upstream activation sequence of Saccharomyces cerevisiae enolase 1 gene. Eur J Biochem. 1989 Sep 15;184(2):305–311. doi: 10.1111/j.1432-1033.1989.tb15020.x. [DOI] [PubMed] [Google Scholar]
- McNeil J. B., Dykshoorn P., Huy J. N., Small S. The DNA-binding protein RAP1 is required for efficient transcriptional activation of the yeast PYK glycolytic gene. Curr Genet. 1990 Dec;18(5):405–412. doi: 10.1007/BF00309909. [DOI] [PubMed] [Google Scholar]
- Morrow B. E., Ju Q., Warner J. R. Purification and characterization of the yeast rDNA binding protein REB1. J Biol Chem. 1990 Dec 5;265(34):20778–20783. [PubMed] [Google Scholar]
- Nishizawa M., Araki R., Teranishi Y. Identification of an upstream activating sequence and an upstream repressible sequence of the pyruvate kinase gene of the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1989 Feb;9(2):442–451. doi: 10.1128/mcb.9.2.442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishizawa M., Suzuki Y., Nogi Y., Matsumoto K., Fukasawa T. Yeast Gal11 protein mediates the transcriptional activation signal of two different transacting factors, Gal4 and general regulatory factor I/repressor/activator site binding protein 1/translation upstream factor. Proc Natl Acad Sci U S A. 1990 Jul;87(14):5373–5377. doi: 10.1073/pnas.87.14.5373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norton I. L., Hartman F. C. Haloacetol phosphates. A comparative study of the active sites of yeast and muscle triose phosphate isomerase. Biochemistry. 1972 Nov 21;11(24):4435–4441. doi: 10.1021/bi00774a003. [DOI] [PubMed] [Google Scholar]
- Pfeifer K., Arcangioli B., Guarente L. Yeast HAP1 activator competes with the factor RC2 for binding to the upstream activation site UAS1 of the CYC1 gene. Cell. 1987 Apr 10;49(1):9–18. doi: 10.1016/0092-8674(87)90750-1. [DOI] [PubMed] [Google Scholar]
- Scott E. W., Allison H. E., Baker H. V. Characterization of TPI gene expression in isogeneic wild-type and gcr1-deletion mutant strains of Saccharomyces cerevisiae. Nucleic Acids Res. 1990 Dec 11;18(23):7099–7107. doi: 10.1093/nar/18.23.7099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tornow J., Santangelo G. M. Efficient expression of the Saccharomyces cerevisiae glycolytic gene ADH1 is dependent upon a cis-acting regulatory element (UASRPG) found initially in genes encoding ribosomal proteins. Gene. 1990 May 31;90(1):79–85. doi: 10.1016/0378-1119(90)90441-s. [DOI] [PubMed] [Google Scholar]
- Uemura H., Fraenkel D. G. gcr2, a new mutation affecting glycolytic gene expression in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Dec;10(12):6389–6396. doi: 10.1128/mcb.10.12.6389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uemura H., Jigami Y. GCR3 encodes an acidic protein that is required for expression of glycolytic genes in Saccharomyces cerevisiae. J Bacteriol. 1992 Sep;174(17):5526–5532. doi: 10.1128/jb.174.17.5526-5532.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uemura H., Jigami Y. Role of GCR2 in transcriptional activation of yeast glycolytic genes. Mol Cell Biol. 1992 Sep;12(9):3834–3842. doi: 10.1128/mcb.12.9.3834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vignais M. L., Sentenac A. Asymmetric DNA bending induced by the yeast multifunctional factor TUF. J Biol Chem. 1989 May 25;264(15):8463–8466. [PubMed] [Google Scholar]