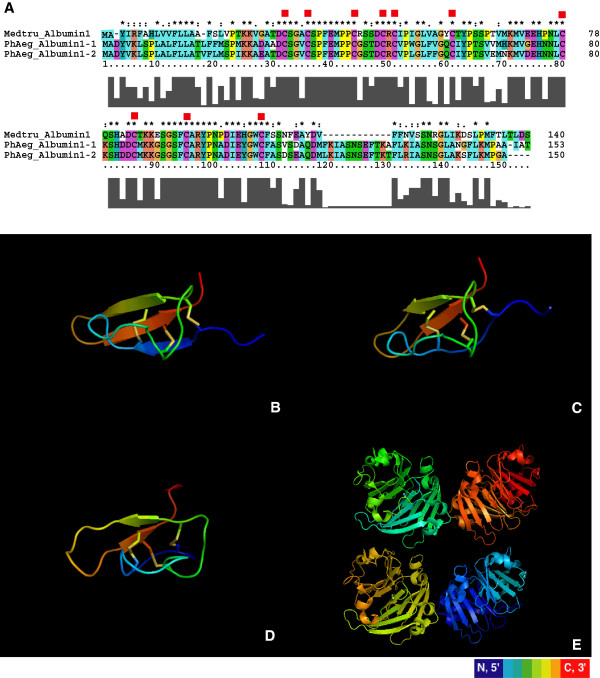
Figure 3.
Amino acid sequence alignment and 3D structure simulation of albumin 1 sequences from Medicago and P. aegyptiaca. (A) Amino acid alignment for the two P. aegyptiaca albumin 1 sequences and a M. truncatula albumin 1 sequence (Q7XZC5, a confirmed KNOTTIN insect toxin protein). Red squares indicate cysteine residues. (B) and (C) show the simulated 3D structures for both Phelipanche sequences. Protein 2D structures are colored from N-terminal to C-terminal with a rainbow color scheme. The three disulfide bonds are shown as colored sticks. The left most and right most sticks open a space that is pierced by the stick in the center. This “disulfide through disulfide knot” is the characteristic structure of KNOTTIN proteins. (D) 3D structure of the KNOTTIN insect toxin protein in M. truncatula. The toxicity of this protein to insect herbivores was confirmed in an earlier report [29]. The PDB file for this 3D structure was obtained from the KNOTTIN database. (E) Predicted albumin 2 (a non-KNOTTIN albumin, PDB ID#3LP9) protein 3D structure in grass pea (Lathyrus sativus).