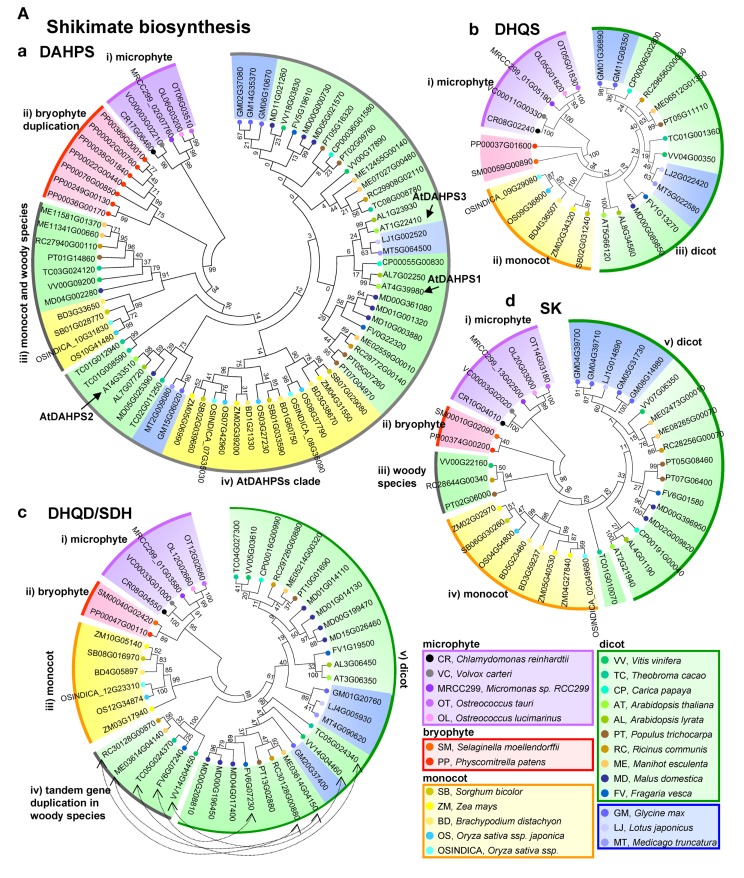
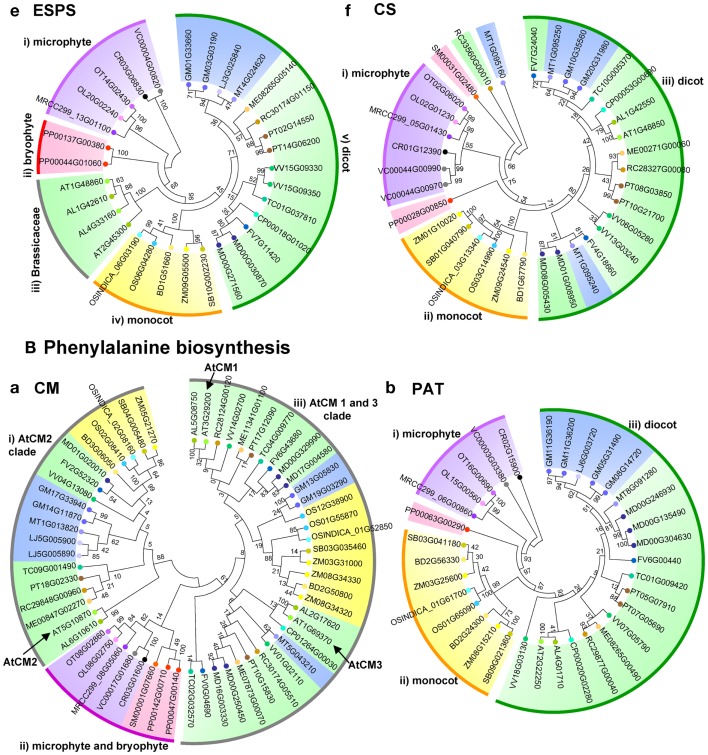
Figure 2.
Phylogenetic tree analysis of shikimate and phenylalanine biosynthetic genes in 25 species. Amino acid sequence phylogenetic trees of (A) shikimate pathway: (a), DAHPS, (b) DHS, (c) DHQD/SD, (d) SK, (e) ESPS, and (f) CS, (B) phenylalanine related genes, (a) CM and (b) PAT. Amino acid sequences of shikimate biosynthetic genes are obtained from Plaza database (http://bioinformatics.psb.ugent.be/plaza/). Relationships among the species considered are presented on the Plaza website. The phylogenetic tree was constructed with the aligned protein sequences by MEGA (version 5.10; http://www.megasoftware.net/; Kumar et al., 2004) using the neighbor-joining method with the following parameters: Poisson correction, complete deletion, and bootstrap (1000 replicates, random seed). The protein sequences were aligned by Plaza. Values on the branches indicate bootstrap support in percentages.