Abstract
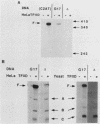
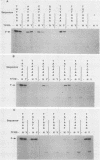
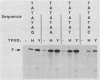
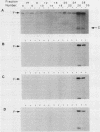
We have analyzed the DNA sequence requirements for TATA element function by assaying the transcriptional activities of 25 promoters, including those representing each of the 18 single-point mutants of the consensus sequence TATAAA, in a reconstituted in vitro system that depends on the TATA element-binding factor TFIID. Interestingly, yeast TFIID and HeLa cell TFIID were virtually identical in terms of their relative activities on this set of promoters. Of the mutated elements, only two had undetectable activity; the rest had activities ranging from 2 to 75% of the activity of the consensus element, which was the most active. In addition, mutations of the nucleotide following the TATAAA core strongly influenced transcriptional activity, although with somewhat different effects on yeast and HeLa TFIID. The activities of all these promoters depended upon TFIID, and the level of TFIID-dependent transcription in vitro correlated strongly with their activities in yeast cells. This suggests that the in vivo activities of these elements reflect their ability to functionally interact with a single TATA-binding factor. However, some elements with similar activities in vitro supported very different levels of transcriptional activation by GAL4 protein in vivo. These results extend the degree of evolutionary conservation between yeast and mammalian TFIID and are useful for predicting the level of TATA element function from the primary sequence.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Breathnach R., Chambon P. Organization and expression of eucaryotic split genes coding for proteins. Annu Rev Biochem. 1981;50:349–383. doi: 10.1146/annurev.bi.50.070181.002025. [DOI] [PubMed] [Google Scholar]
- Bucher P., Trifonov E. N. Compilation and analysis of eukaryotic POL II promoter sequences. Nucleic Acids Res. 1986 Dec 22;14(24):10009–10026. doi: 10.1093/nar/14.24.10009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buratowski S., Hahn S., Guarente L., Sharp P. A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell. 1989 Feb 24;56(4):549–561. doi: 10.1016/0092-8674(89)90578-3. [DOI] [PubMed] [Google Scholar]
- Buratowski S., Hahn S., Sharp P. A., Guarente L. Function of a yeast TATA element-binding protein in a mammalian transcription system. Nature. 1988 Jul 7;334(6177):37–42. doi: 10.1038/334037a0. [DOI] [PubMed] [Google Scholar]
- Cavallini B., Faus I., Matthes H., Chipoulet J. M., Winsor B., Egly J. M., Chambon P. Cloning of the gene encoding the yeast protein BTF1Y, which can substitute for the human TATA box-binding factor. Proc Natl Acad Sci U S A. 1989 Dec;86(24):9803–9807. doi: 10.1073/pnas.86.24.9803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavallini B., Huet J., Plassat J. L., Sentenac A., Egly J. M., Chambon P. A yeast activity can substitute for the HeLa cell TATA box factor. Nature. 1988 Jul 7;334(6177):77–80. doi: 10.1038/334077a0. [DOI] [PubMed] [Google Scholar]
- Chen W., Struhl K. Saturation mutagenesis of a yeast his3 "TATA element": genetic evidence for a specific TATA-binding protein. Proc Natl Acad Sci U S A. 1988 Apr;85(8):2691–2695. doi: 10.1073/pnas.85.8.2691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen W., Struhl K. Yeast upstream activator protein GCN4 can stimulate transcription when its binding site replaces the TATA element. EMBO J. 1989 Jan;8(1):261–268. doi: 10.1002/j.1460-2075.1989.tb03372.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Concino M. F., Lee R. F., Merryweather J. P., Weinmann R. The adenovirus major late promoter TATA box and initiation site are both necessary for transcription in vitro. Nucleic Acids Res. 1984 Oct 11;12(19):7423–7433. doi: 10.1093/nar/12.19.7423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davison B. L., Egly J. M., Mulvihill E. R., Chambon P. Formation of stable preinitiation complexes between eukaryotic class B transcription factors and promoter sequences. Nature. 1983 Feb 24;301(5902):680–686. doi: 10.1038/301680a0. [DOI] [PubMed] [Google Scholar]
- Eisenmann D. M., Dollard C., Winston F. SPT15, the gene encoding the yeast TATA binding factor TFIID, is required for normal transcription initiation in vivo. Cell. 1989 Sep 22;58(6):1183–1191. doi: 10.1016/0092-8674(89)90516-3. [DOI] [PubMed] [Google Scholar]
- Giangrande A., Mettling C., Martin M., Ruiz C., Richards G. Drosophila Sgs3 TATA: effects of point mutations on expression in vivo and protein binding in vitro with staged nuclear extracts. EMBO J. 1989 Nov;8(11):3459–3466. doi: 10.1002/j.1460-2075.1989.tb08510.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gill G., Ptashne M. Negative effect of the transcriptional activator GAL4. Nature. 1988 Aug 25;334(6184):721–724. doi: 10.1038/334721a0. [DOI] [PubMed] [Google Scholar]
- Hahn S., Buratowski S., Sharp P. A., Guarente L. Isolation of the gene encoding the yeast TATA binding protein TFIID: a gene identical to the SPT15 suppressor of Ty element insertions. Cell. 1989 Sep 22;58(6):1173–1181. doi: 10.1016/0092-8674(89)90515-1. [DOI] [PubMed] [Google Scholar]
- Hahn S., Buratowski S., Sharp P. A., Guarente L. Yeast TATA-binding protein TFIID binds to TATA elements with both consensus and nonconsensus DNA sequences. Proc Natl Acad Sci U S A. 1989 Aug;86(15):5718–5722. doi: 10.1073/pnas.86.15.5718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harbury P. A., Struhl K. Functional distinctions between yeast TATA elements. Mol Cell Biol. 1989 Dec;9(12):5298–5304. doi: 10.1128/mcb.9.12.5298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helmann J. D., Chamberlin M. J. Structure and function of bacterial sigma factors. Annu Rev Biochem. 1988;57:839–872. doi: 10.1146/annurev.bi.57.070188.004203. [DOI] [PubMed] [Google Scholar]
- Hobson G. M., Mitchell M. T., Molloy G. R., Pearson M. L., Benfield P. A. Identification of a novel TA-rich DNA binding protein that recognizes a TATA sequence within the brain creatine kinase promoter. Nucleic Acids Res. 1988 Sep 26;16(18):8925–8944. doi: 10.1093/nar/16.18.8925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodo H. G., 3rd, Blatti S. P. Purification using polyethylenimine precipitation and low molecular weight subunit analyses of calf thymus and wheat germ DNA-dependent RNA polymerase II. Biochemistry. 1977 May 31;16(11):2334–2343. doi: 10.1021/bi00630a005. [DOI] [PubMed] [Google Scholar]
- Homa F. L., Glorioso J. C., Levine M. A specific 15-bp TATA box promoter element is required for expression of a herpes simplex virus type 1 late gene. Genes Dev. 1988 Jan;2(1):40–53. doi: 10.1101/gad.2.1.40. [DOI] [PubMed] [Google Scholar]
- Horikoshi M., Wang C. K., Fujii H., Cromlish J. A., Weil P. A., Roeder R. G. Cloning and structure of a yeast gene encoding a general transcription initiation factor TFIID that binds to the TATA box. Nature. 1989 Sep 28;341(6240):299–303. doi: 10.1038/341299a0. [DOI] [PubMed] [Google Scholar]
- Huang D. H., Horikoshi M., Roeder R. G. Activation of the adenovirus EIIa late promoter by a single-point mutation which enhances binding of transcription factor IID. J Biol Chem. 1988 Sep 5;263(25):12596–12601. [PubMed] [Google Scholar]
- Lobo S. M., Hernandez N. A 7 bp mutation converts a human RNA polymerase II snRNA promoter into an RNA polymerase III promoter. Cell. 1989 Jul 14;58(1):55–67. doi: 10.1016/0092-8674(89)90402-9. [DOI] [PubMed] [Google Scholar]
- Matsui T., Segall J., Weil P. A., Roeder R. G. Multiple factors required for accurate initiation of transcription by purified RNA polymerase II. J Biol Chem. 1980 Dec 25;255(24):11992–11996. [PubMed] [Google Scholar]
- Myers R. M., Tilly K., Maniatis T. Fine structure genetic analysis of a beta-globin promoter. Science. 1986 May 2;232(4750):613–618. doi: 10.1126/science.3457470. [DOI] [PubMed] [Google Scholar]
- Nakajima N., Horikoshi M., Roeder R. G. Factors involved in specific transcription by mammalian RNA polymerase II: purification, genetic specificity, and TATA box-promoter interactions of TFIID. Mol Cell Biol. 1988 Oct;8(10):4028–4040. doi: 10.1128/mcb.8.10.4028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker C. S., Topol J. A Drosophila RNA polymerase II transcription factor contains a promoter-region-specific DNA-binding activity. Cell. 1984 Feb;36(2):357–369. doi: 10.1016/0092-8674(84)90229-0. [DOI] [PubMed] [Google Scholar]
- Ponticelli A. S., Struhl K. Analysis of Saccharomyces cerevisiae his3 transcription in vitro: biochemical support for multiple mechanisms of transcription. Mol Cell Biol. 1990 Jun;10(6):2832–2839. doi: 10.1128/mcb.10.6.2832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reinberg D., Horikoshi M., Roeder R. G. Factors involved in specific transcription in mammalian RNA polymerase II. Functional analysis of initiation factors IIA and IID and identification of a new factor operating at sequences downstream of the initiation site. J Biol Chem. 1987 Mar 5;262(7):3322–3330. [PubMed] [Google Scholar]
- Reinberg D., Roeder R. G. Factors involved in specific transcription by mammalian RNA polymerase II. Purification and functional analysis of initiation factors IIB and IIE. J Biol Chem. 1987 Mar 5;262(7):3310–3321. [PubMed] [Google Scholar]
- Samuels M., Fire A., Sharp P. A. Separation and characterization of factors mediating accurate transcription by RNA polymerase II. J Biol Chem. 1982 Dec 10;257(23):14419–14427. [PubMed] [Google Scholar]
- Sawadogo M., Roeder R. G. Factors involved in specific transcription by human RNA polymerase II: analysis by a rapid and quantitative in vitro assay. Proc Natl Acad Sci U S A. 1985 Jul;82(13):4394–4398. doi: 10.1073/pnas.82.13.4394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt M. C., Kao C. C., Pei R., Berk A. J. Yeast TATA-box transcription factor gene. Proc Natl Acad Sci U S A. 1989 Oct;86(20):7785–7789. doi: 10.1073/pnas.86.20.7785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shapiro D. J., Sharp P. A., Wahli W. W., Keller M. J. A high-efficiency HeLa cell nuclear transcription extract. DNA. 1988 Jan-Feb;7(1):47–55. doi: 10.1089/dna.1988.7.47. [DOI] [PubMed] [Google Scholar]
- Simon M. C., Fisch T. M., Benecke B. J., Nevins J. R., Heintz N. Definition of multiple, functionally distinct TATA elements, one of which is a target in the hsp70 promoter for E1A regulation. Cell. 1988 Mar 11;52(5):723–729. doi: 10.1016/0092-8674(88)90410-2. [DOI] [PubMed] [Google Scholar]
- Singer V. L., Wobbe C. R., Struhl K. A wide variety of DNA sequences can functionally replace a yeast TATA element for transcriptional activation. Genes Dev. 1990 Apr;4(4):636–645. doi: 10.1101/gad.4.4.636. [DOI] [PubMed] [Google Scholar]
- Smale S. T., Baltimore D. The "initiator" as a transcription control element. Cell. 1989 Apr 7;57(1):103–113. doi: 10.1016/0092-8674(89)90176-1. [DOI] [PubMed] [Google Scholar]
- Struhl K. Constitutive and inducible Saccharomyces cerevisiae promoters: evidence for two distinct molecular mechanisms. Mol Cell Biol. 1986 Nov;6(11):3847–3853. doi: 10.1128/mcb.6.11.3847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Struhl K. Yeast HIS3 expression in Escherichia coli depends upon fortuitous homology between eukaryotic and prokaryotic promoter elements. J Mol Biol. 1986 Sep 20;191(2):221–229. doi: 10.1016/0022-2836(86)90259-7. [DOI] [PubMed] [Google Scholar]