Figure 1.
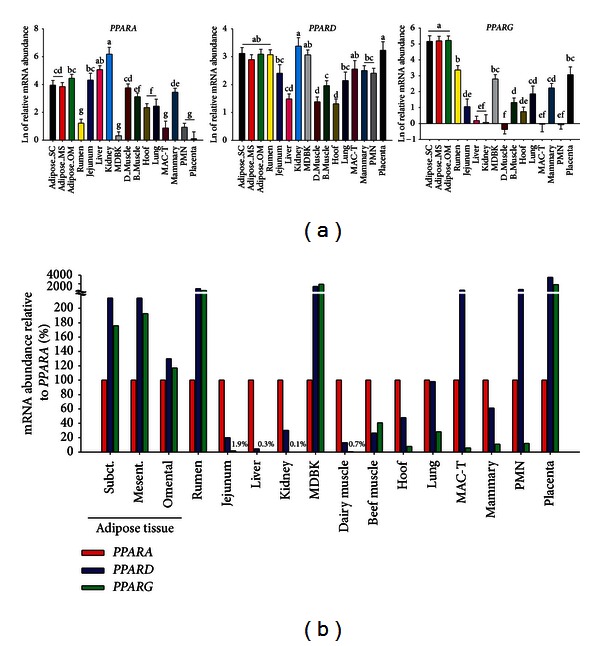
(a) Relative transcript abundance of each PPAR isotype in several bovine tissues and cells. We measured gene expression of PPAR isotypes in 14 different tissues including tissues from adult dairy cattle: adipose tissue (subcutaneous, mesenteric, and omental), small intestine (jejunum), liver, hoof corium, lung, kidney, mammary gland, blood polymorphonuclear leukocytes (PMN), and placenta; from dairy calves: rumen papillae and semitendinosus muscle (D-muscle); skeletal muscle of beef cattle (Longissimus lombarum); and two cell lines: Madin-Darby Bovine Kidney (MDBK) and bovine mammary alveolar cells (MAC-T). The total RNA was extracted and qPCR performed as previously described [26]. The qPCR data were normalized by the geometrical mean of 5 internal control genes (PPP1R11, RPS15A, ACTB1, MRPL39, and UXT). For the difference of each PPAR isotype abundance between tissues, the qPCR data were transformed using a 6-point standard curve prior statistical analysis using PROC GLM of SAS (version 9.3) with tissue as main effect. Dissimilar letters denote significant differences (P < 0.05). (b) Tissue-specific relative mRNA abundance between PPAR isotypes. The % relative abundance of the three PPAR isotypes in each tissue was calculated using the delta Ct method as previously described [27]. The final data for PPARG and PPARD were obtained as % relative to PPARA. N.B.: the y-axis values in (a) are least square means of the Ct values transformed using the standard curve and then log2-transformed. The values in (b) are calculated without use of a standard curve. Therefore, the values in (a) are radically different compared to the values in (b) and the two cannot be compared.
