Figure 4.
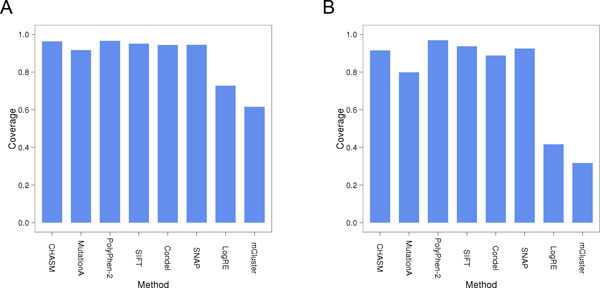
Coverage of prediction. CHASM, MutationA(ssessor), PolyPhen-2, SIFT, Condel, SNAP, and CHASM scored most missense mutations. LogRE and mCluster predictions are restricted to alterations that occur in domain regions and so scored less than 80% of likely cancer drivers (A). Coverage of likely-neutral mutations (B) was broadly similar, but with even lower coverage for LogRE and mCluster due to the lower prevalence of neutral mutations in domains.
