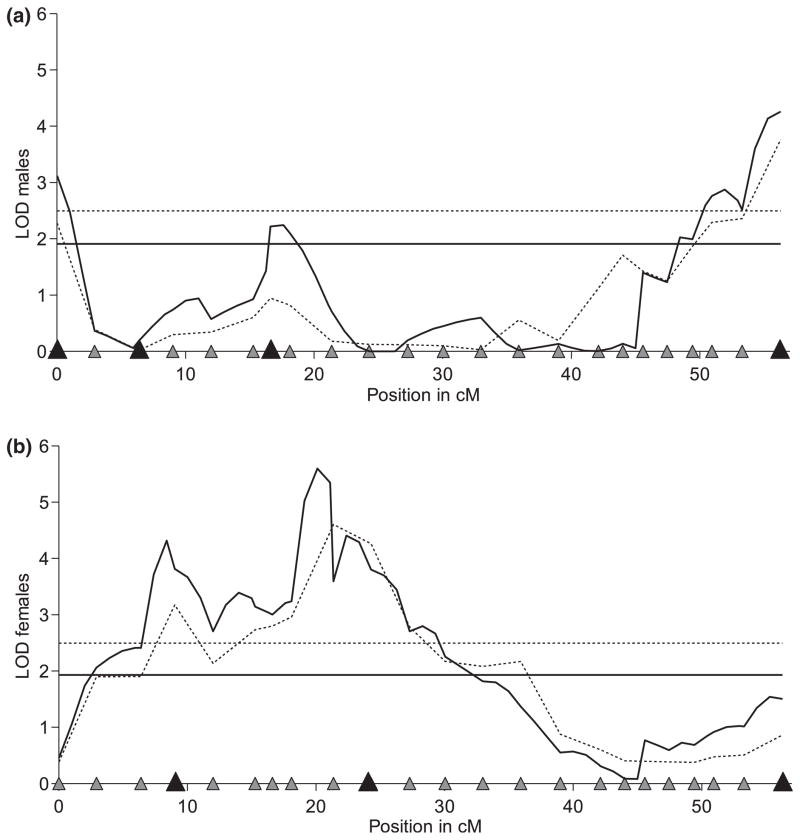
Fig. 3.
Results of interval-mapping methods for ln[chill coma recovery time (CCRT)] in males (a) and females (b) of Drosophila melanogaster. The one-quantitative trait loci (QTL) analysis (dotted lines) and the composite interval mapping (CIM) (solid lines) for males and females give consistent results, but the CIM better resolved the genetic architecture of multiple QTL for cold tolerance on the X chromosome. The horizontal lines correspond to the threshold values (dotted line for the one-QTL model and the solid one for the CIM). The distribution of the 23 genetic markers used is shown on the x-axis by triangles, where larger black triangles indicate the locations of QTL cofactors. Each position at which the LOD score exceeds the threshold value is a genomic region showing significant association with CCRT—i.e. a QTL.