Table 2. Data used to construct both the initial and gold-standard networks used in the evaluation of PANDA and the other network reconstruction algorithms.
| Network Name | Data used to construct network[reference] | Number of TFs/Genes/Conditions |
Initial Cooperativity Network (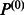 ) ) |
Affinity Capture-MS [47], [48] | TFs: 53 (43 with evidence) |
Gold Standard Cooperativity Network (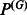 ) ) |
Low-throughput evidence [47], [48] | TFs: 33 |
Initial Regulatory Network (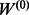 ) ) |
Motif [45], [46] | TFs: 53 Genes: 2555 |
Gold Standard Regulatory Network (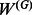 ) ) |
ChIP-chip [45], [49] | TFs: 52 Genes: 1073 |
Initial Co-regulatory Network – Knock-Out Data (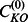 ) ) |
Gene Expression [43], [44] | Genes: 2555 Conditions: 106 |
Gold Standard Co-regulatory Network (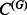 ) ) |
ChIP-chip [45], [49] | Genes:1073 (1072 co-targeted) |
Initial Co-regulatory Network – Cell Cycle Data (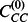 ) ) |
Gene Expression [51]–[54] | Genes: 2555 Conditions: 56 |
Initial Co-regulatory Network – Stress Response Data (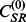 ) ) |
Gene Expression [55], [56] | Genes: 2555 Conditions:173 |
References include both the publication and the website from which the normalized data was downloaded. The number of transcription factors and genes reported are those used to construct each network. The 53 transcription factors and 2555 genes mentioned in the initial protein-cooperativity network, initial regulatory network and initial co-regulatory networks are the same set of TFs and genes for all networks, and represent those for which we had both motif and expression data. For the initial cooperativity network, we allowed TFs for which we had no Affinity Capture-MS data to be initialized as self-cooperating (Pii = 1). The transcription factors and genes used to construct the three gold-standard networks are the subset of the aforementioned 53 TFs and 2555 genes for which we had ChIP information (52 TFs and 1073 genes), or “low-throughput” interaction information (33 TFs). We used this subset of TFs and genes when evaluating the quality of each network. “Low-throughput evidence” data represents interactions with evidence from “co-fractionation”, “co-localization”, “FRET” or “reconstituted complex.”
