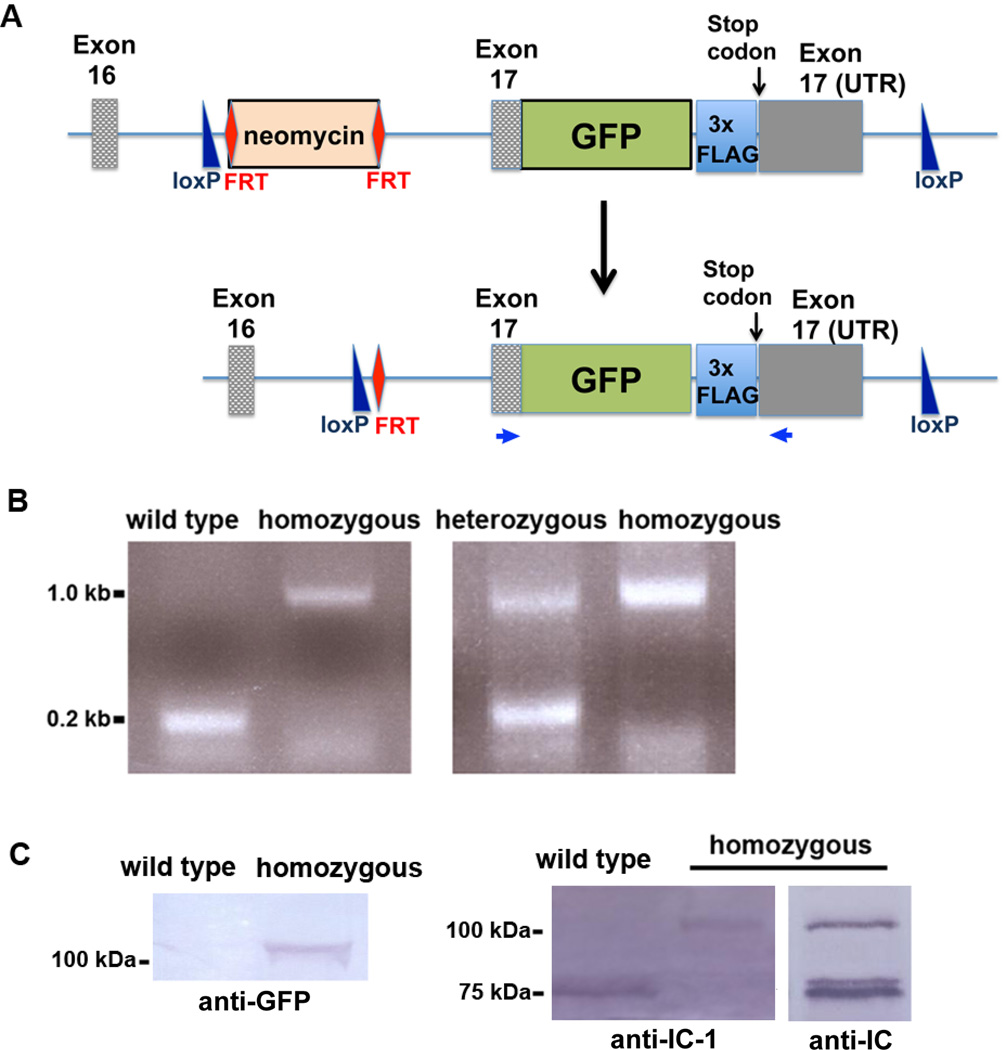
Figure 1.
Construction and characterization of the dynein IC-1 knock-in mice. (A) A diagram showing the in-frame insertion of the GFP tag and the 3xFLAG tag before the stop codon of the endogenous Dync1i1 gene. The selection marker, the FRT-site-flanked neomycin gene, was inserted in the intron between exons 16 and 17. Note that the coding region of exon 17 is labeled as “Exon 17” and the 3’ untranslated region (UTR) of exon 17 is labeled as “Exon 17 (UTR)”. Two loxP sites were also inserted to flank exon 17, which would potentially aid in the deletion of the coding region of exon 17 and its 3’ untranslated region for making a partial IC-1 knockout mouse model. The neomycin marker was removed by breeding with Flp delete mice. The two primers used for the PCR analysis shown in B are indicated as small blue arrows. (B) A PCR-based method to screen for homozygous and heterozygous knock-in mice. Positions of the primers are shown in A. This pair of primers generates a 0.2 kb product for the wild type, a 1 kb product for the homozygous knock-in mice and both of these products for the heterozygous knock-in mice. (C) Western blots showing the IC-1-GFP-3xFLAG fusion protein of ~100 kDa recognized by an anti-GFP antibody, the IC-1-specific antibody (labeled as “anti-IC-1”) and the general anti-IC antibody 74.1 (labeled as “anti-IC”). Note that the anti-IC-1 antibody only recognized the ~75 kDa IC-1 protein in the wild type sample and the ~100 kDa fusion protein in the homozygous sample while the anti-IC antibody recognized both the IC-1-GFP-3xFLAG fusion protein (~100 kDa) and the IC-2 protein (~75 kDa) in the homozygous sample.