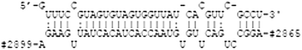Table 3.
The predicted interactors to the two tRF-5s.
| tRF | Gene name [accession number] | Base-paring to potential target sites | ΔG value (Kcal/mol) | |
|---|---|---|---|---|
|
tRF5-ValGTG |
Homo sapiens protein kinase C, beta (PRKCB), transcript variant 1, mRNA [NM_121535.2] |
|
-34.2 |
|
| Homo sapiens carboxyl ester lipase (bile salt-stimulated lipase) (CEL), mRNA [NM_001807.3] |
|
-35.3 |
||
| HOMO sapiens glutamyl-tRNA synthetase 2, mitochondrial (EARS2), transcript variant 1, mRNA [NM_001083614.1] |
|
-40.2 |
||
| HOMO sapiens solute carrier family 23 (nucleobase trnasporters), member 2 (SLC23A2), trnascript variant 1, mRNA [NM_005116.5] |
|
-36.9 |
||
| tRF5-GlyGCC | Homo sapiens long interegenic non-protein coding RNA 317 (LINC00317), non-coding RNA [NR_038872.1] |
|
-50.6 |
|
| Homo sapiens SH3-domain GRB2-like endophilin B1 (SH3GLB1), transcript variant 1, mRNA [NM_016009.4] |
|
-37.3 |
||
| Homo sapiens glial cells missing homolog 1 (drosophila) (GMC1), mRNA [NM_016009.4] |
|
-37.2 |
||
| Homo sapiens syntrophi, beta 1 (dystrophin-asociated protein A1, 59kDa, basic component 1) (SNTB1) mRNA [NM_021021.3 | -39.7 |
The potential target mRNAs of the two tRF-5s and their interactions are summarized and tabulated. In the depicted base-pairing interactions, the sequences on the top and bottom are tRF-5 and its predicted target mRNA, respectively. The numbers of target mRNAs indicate nucleotide coordinates of the RefSeq shown in the second column. The degree of interaction, expressed in ∆ G values, was calculated by the RNAhybrid program and shown.