Table 2.
Mapping Results.
| Species | Method | Total RNA (µg) | Reads | Mapped | Percentage | Filtereda | Percentage |
|---|---|---|---|---|---|---|---|
| mel | CAGE | 5 | 858,863 | 372,550 | 43 | 166,096 | 19b |
| mel | CAGE | 10 | 119,426 | 58,556 | 49 | NA | NA |
| sim | CAGE | 5 | 1,180,578 | 823,516 | 70 | 320,263 | 27 |
| sec | CAGE | 5 | 1,003,687 | 695,863 | 69 | 287,079 | 29 |
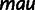 c c
|
CAGE | 5 | 627,904 | 286,214 | 46 | NA | NA |
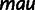 |
CAGE | 5 | 1,201,745 | 488,569 | 41 | NA | NA |
| pse | CAGE | 5 | 1,161,039 | 822,188 | 71 | 323,942 | 28 |
| mel | RNAseq | 5 | 1,209,363 | 706,386 | 58 | 1,232,707 | 43b |
| mel | RNAseq | 10 | 1,664,075 | 1,004,598 | 60 | NA | NA |
Note.—NA, note applicable.
aMapped reads after removing PCR duplicates.
b5 μg and 10 μg samples were pooled.
cDrosophila mauritiana was used for reproducibility estimates only.
