Table 1. Network parameters of the biomolecular regulatory networks.
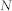 |
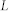 |
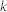 |
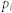 |
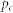 |
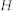 |
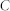 |
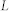 |
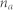 |
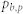 |
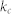 |
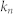 |
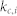 |
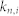 |
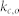 |
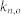 |
|
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| S. cerevisiae cell cycle network | 11 | 34 | 6.18 | 0.56 | 0.36 | 0.38 | 0.277 | 2.12 | 7 | 0.86 | 4.5 | 7.14 | 3.25 | 2.29 | 1.25 | 3.43 |
| S. pombe cell cycle network | 9 | 26 | 5.78 | 0.69 | 0.44 | 0.27 | 0 | 1.984 | 16 | 0.67 | 4.5 | 6.8 | 3 | 2 | 1.5 | 2.33 |
| GRN underlying mammalian cortical area development | 10 | 19 | 3.8 | 0.47 | 0.1 | 0.26 | 0 | 2.711 | 3 | 0.77 | 4.33 | 3 | 2 | 1.75 | 3.85 | 1.25 |
| GRN underlying A. thaliana development | 15 | 41 | 5.47 | 0.42 | 0.067 | 0.67 | 0.287 | 2.189 | 3 | 0.71 | 8.67 | 5.89 | 3 | 3.11 | 5.67 | 1.89 |
| GRN underlying mouse myeloid development | 11 | 30 | 5.45 | 0.47 | 0.27 | 0.43 | 0.439 | 2.081 | 8 | 0.53 | 6.33 | 5.13 | 2.33 | 2.38 | 3.33 | 2 |
| Mammalian cell cycle network | 20 | 51 | 5.1 | 0.24 | 0.05 | 0.43 | 0.112 | 2.209 | NA | 0.998 | 6.5 | 5.06 | 4 | 2.53 | 2.5 | 2.53 |
| CREB signaling network | 64 | 152 | 4.75 | 0.22 | 0.078 | 0.75 | 0.051 | 3.878 | NA | 0.29 | 7.5 | 4.59 | 4.5 | 2.37 | 3 | 2.22 |
| Human fibroblast signaling network | 139 | 575 | 8.27 | 0.23 | 0.086 | 0.71 | 0.104 | 4.486 | NA | 0.56 | 9.67 | 8.62 | 3.5 | 4.07 | 4.17 | 3.86 |
 , number of nodes;
, number of nodes;  , number of links;
, number of links;  , average degree;
, average degree; 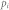 , proportion of inhibitory links;
, proportion of inhibitory links; 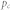 , proportion of control kernel nodes;
, proportion of control kernel nodes;  degree heterogeneity;
degree heterogeneity;  , clustering coefficient;
, clustering coefficient;  , characteristic path length;
, characteristic path length; 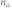 , number of attractors;
, number of attractors; 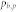 , proportion of the basin size of the primary attractor;
, proportion of the basin size of the primary attractor; 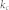 average degree of the control kernel nodes;
average degree of the control kernel nodes; 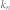 , average degree of the non-kernel nodes;
, average degree of the non-kernel nodes; 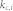 , average in-degree of the control kernel nodes;
, average in-degree of the control kernel nodes; 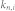 , average in-degree of the non-kernel nodes;
, average in-degree of the non-kernel nodes; 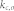 , average out-degree of the control kernel nodes;
, average out-degree of the control kernel nodes; 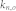 , average out-degree of the non-kernel nodes; NA, not available. The exact numbers of attractors for the mammalian cell cycle network, the CREB signaling network, and the human fibroblast signaling network could not be obtained because there were large numbers of attractors with small basins.
, average out-degree of the non-kernel nodes; NA, not available. The exact numbers of attractors for the mammalian cell cycle network, the CREB signaling network, and the human fibroblast signaling network could not be obtained because there were large numbers of attractors with small basins.
