Figure 9.
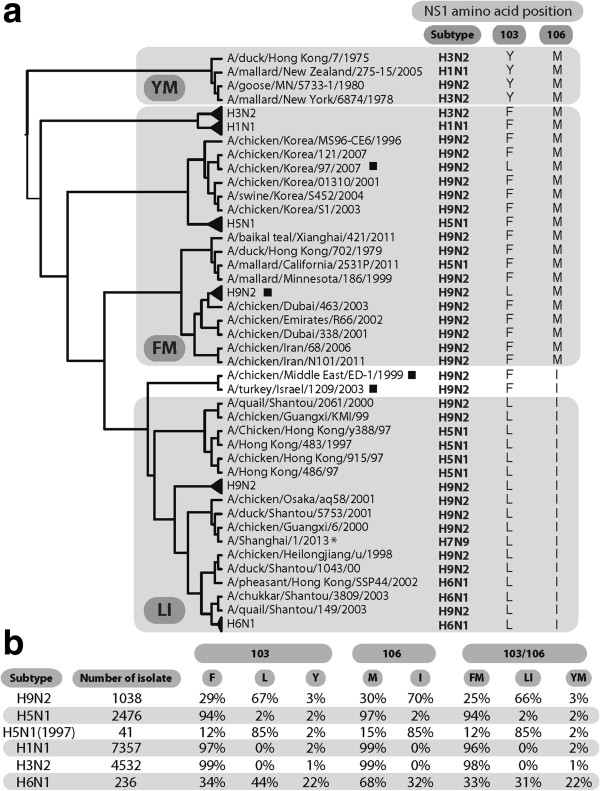
Analysis of NS1 protein 103 and 106 residue mutation in natural human and animal isolates. a. Phylogenetic trees based on the NS1 protein sequence of influenza A viruses isolated from subtypes H1N1, H3N2, H5N1, H6N1and H9N2. Know subtypes and amino acid at position 103 and 106 are indicated. Different 103/106 residue motifs (FM, YM, LI) are represented by a grey square. Full trees and a list of GenBank accession numbers of the viruses used are provided in supplementary information. Independent selection of F103L occurred in the FM lineage and are indicated with a square (■) and independent selection of M106I is shown for 2 H9N2 isolates representative of a Middle Eastern lineage. b. Variability found at position 103 and 106 in different IAV subtypes. H1N1, H3N2, H5N1, H6N1 and H9N2 viruses were analyzed for the frequency of different amino acid compositions at position 103, 106 and 103/106.