Figure 1.
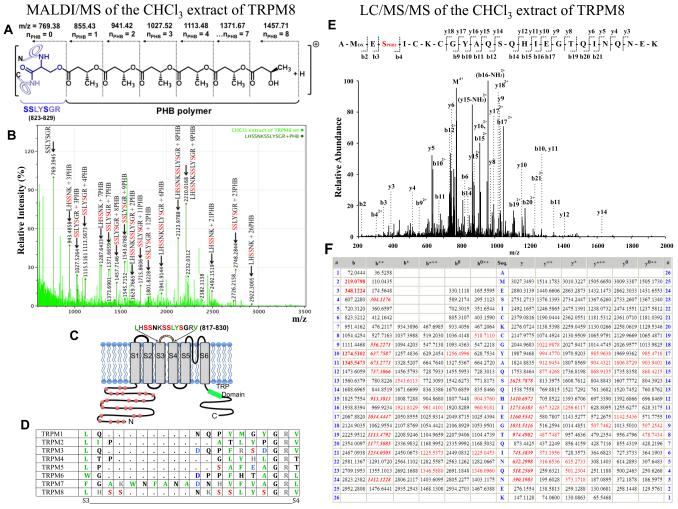
Mass spectrometric analysis of the chloroform-extracted peptide LHSSNKSSLYSGR (817–829) of the TRPM8 protein derived from MALDI/MS experiments:
Panel A: Molecular composition of PHBylated serine residue on the SSLYSGR (823–829) peptide with a number of PHB units attached via an ester bond, numbers indicate the PHB modification with a shift in the monoisotopic masses.
Panel B: Intensity peaks of the LHSSNKSSLYSGR (817–829) peptide with indicated PHB modifications detected in mass spectrum of the chloroform-extracted peptides derived from MALDI/MS. Masses were analyzed with the ExPASy FindMod Tool (Swiss Proteomics Bioinformatics Resources) run against the TRPM8 sequence with possible PHB modifications up to 30 units, with 0 or 1 missed cleavage cites for trypsin-digested protein (error window ±50 ppm). The variability of different length of PHB might be due to the breakage of labile ester bonds under the MS laser beam (Xian et al., 2007).
Panel C: Cartoon of the putative PHB-modification sites on the TRPM8 protein with a sequence indication for the extracellular PHBylated peptides. The red spheres indicate putative PHBylated peptides on the N-terminus of TRPM8, derived from MALDI-MS experiments. D: The amino-acid sequence of the S3–S4 linker of TRPM family ion channels is not conserved.
Panel E: MS/MS spectrum of the quadruply charged ion (m/z 775.354) corresponding to the peptide 63AMESICKCGYAQSQHIEGTQINQNEK88 showing the methionine oxidation and 2 units of PHB modification on serine residues. The observed y- and b-ion series confirmed the peptide sequence. The b4, b9, b10, b11, b12, b14, b15, b16, b17, b19, b20, and b21 ions confirmed PHB localization on the serine residue.
Panel F: A table showing the identified peptide fragment ions from the spectrum (red) versus theoretical fragment ions not found in the spectrum (black). The bold italic red ions are the relatively abundant y- or b-ions that contributed to the scoring of the peptide and PTM identification. Additional matched ions, including bold red ions, were not used for the calculation of the identification score.