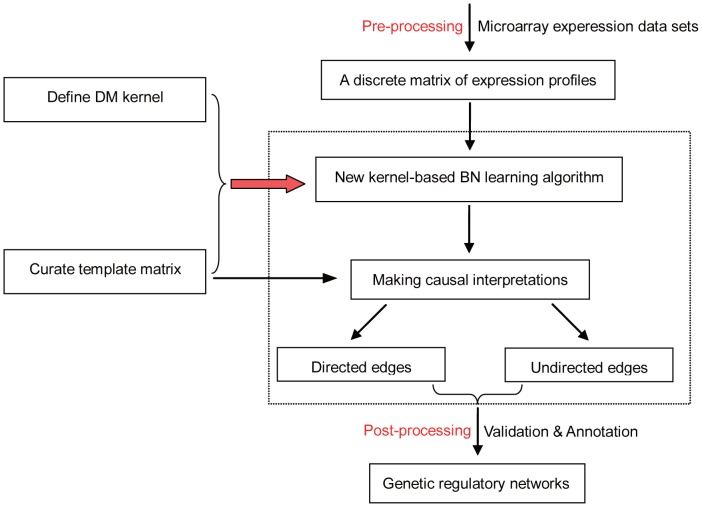
Figure 1. Overview of the Bayesian network learning algorithm DM_BN for reverse engineering regulatory networks from genetic perturbation data.
As an example, DM_BN takes as input deletion mutant gene expression profiles. The relative change of the mRNA expression levels of the deletion mutant strains versus the wild type (WT) is represented by 1 (significant up-regulation), −1 (significant down-regulation) and 0 (no significant change). The algorithm incorporate a new kernel to model the consistent gene expression changes upon perturbation and it employs a template of all potential regulator-regulator interactions to enable more accurate and much faster BN learning. After learning the BN structure, Meek's rule [28] is used to infer compelled (directed) and non-compelled (undirected) edges. The precision of the inferred network is then assessed by existing knowledge of protein complexes, regulatory relationships (MAPK signaling transduction pathways) and gene annotations.