Figure 3. DNA blot analysis of maize transgenic events.
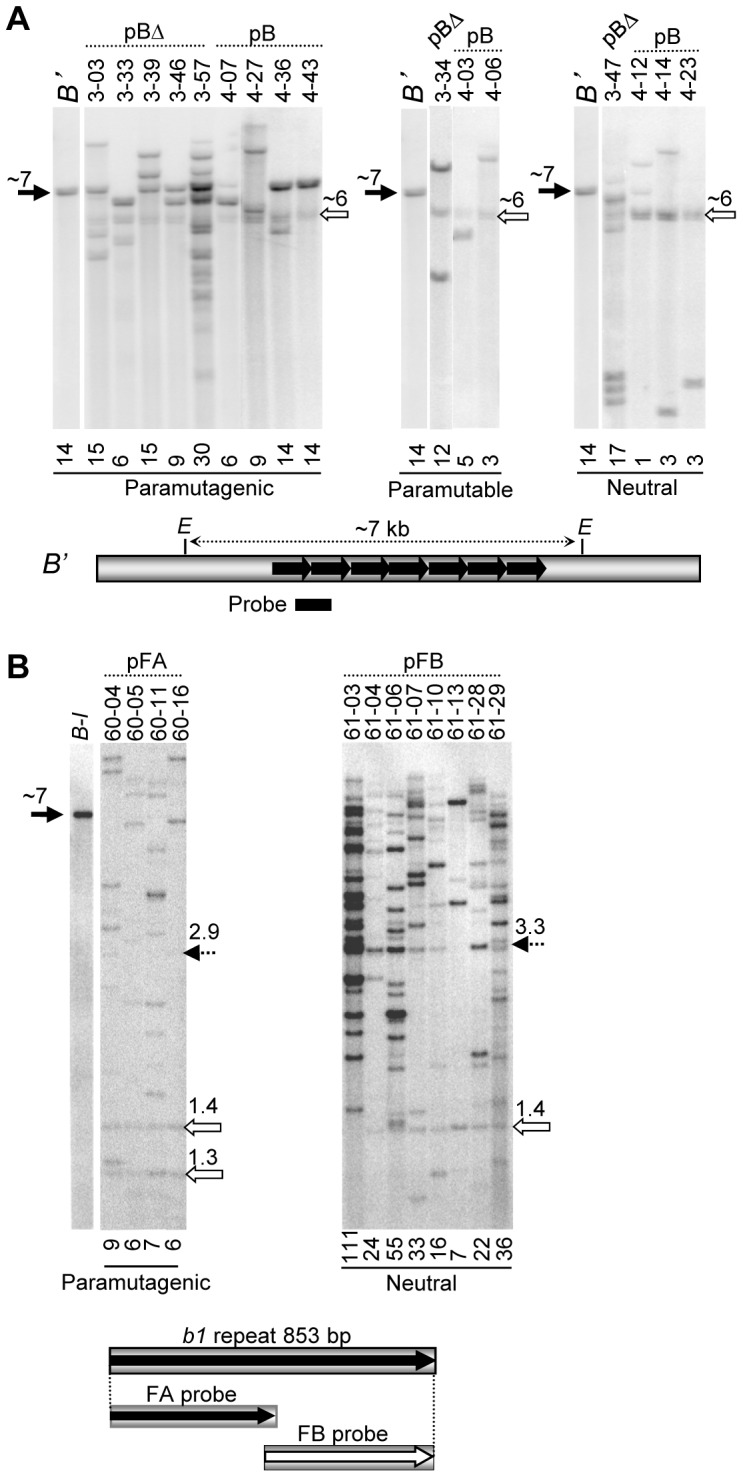
The numbers above the arrows indicate approximate fragment sizes in kb. A. DNA blot analysis of the paramutagenic, paramutable, and neutral pB and pBΔ transgenic loci. Genomic DNA from transgenic plants was digested with EcoRI (E) and blots were hybridized with the tandem repeat probe shown as a bar below the map. The B' allele was used as a control to indicate the ∼7 kb EcoRI fragment containing the seven tandem repeats (black arrow). All transgenic plants were heterozygous for two different neutral b1 alleles, each containing a single copy of the repeat unit, together producing a ∼6 kb doublet upon digestion (open arrow). Transgenic event number and construct names are shown above the lanes, while the approximate repeat copy number, estimated using phosphor imaging analysis, is shown below each lane. B. DNA blot analyses of plants carrying the pFA or pFB transgenes; genomic DNA was digested with BamHI and BglII, which cut on either side of the tandem repeat array. The FA and FB fragments diagrammed below the blots were used as probes. Bands corresponding to B-I (∼7 kb) and the neutral b1 alleles (1.4 kb and 1.3 kb) are indicated by black and open arrows, respectively. The 2.9 and 3.3 kb bands corresponding to the intact FA and FB tandem arrays, respectively, are indicated by dotted arrows.
