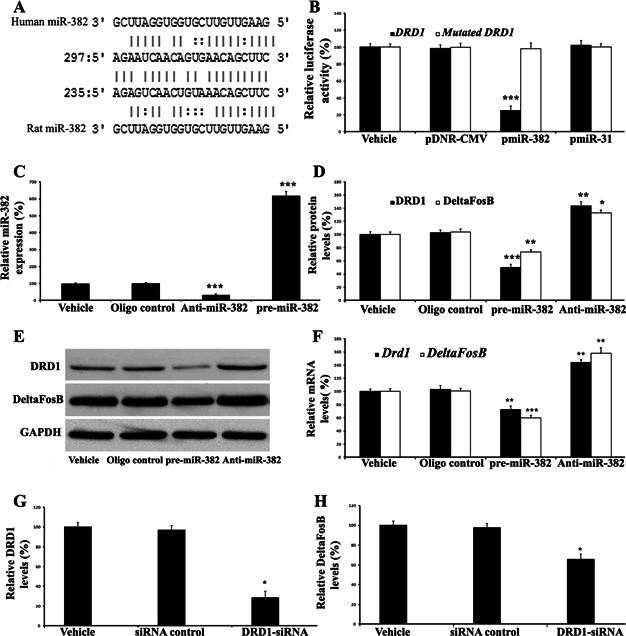
Figure 3. Drd1 is a direct target gene of miR-382 and is a regulator for the expression of DeltaFosB in cultured CAD cells: **p < 0.01, ***p < 0.001, Student's t-test.
Source data is available for this figure in the Supporting Information.
- Drd1 is a potential target gene of miR-382 predicted by computational analysis.
- The luciferase reporter construct, containing the putative miR-382 binding sequence from 3′-UTR of rat Drd1 gene, was transfected into HEK293 cells with vehicle (Vehicle), an empty vector (pDNR-CMV), miR-382 (pmiR-31) or a control plasmid expressing an unrelated miRNA, miR-31 (pmiR-31). The construct with mutated fragment of the 3′-UTR of Drd1 mRNA without the putative miR-382 binding sequences was used as the mutated control (mutated Drd1), pmiR-382, but not pmiR-31 or pDNR-CMV, inhibited luciferase activity (p = 1.17571599413E-6). Values are mean ± SEM from 6 independent experiments (n = 6), compared with that in control (pDNR-CMV). In the mutated control group, the inhibitory effect of pmiR-miR-382 on luciferase activity disappeared. Values are mean ± SEM from 3 independent experiments (n = 3), compared with that in control (pDNR-CMV).
- The expression of miR-382 in cultured CAD cells was modulated by LNA-anti-miR-382 (Anti-miR-382) and pre-miR-382. Vehicle and control oligos (oligo control) were used as controls. miR-382 expression was down-regulated by LNA-anti-miR-382 (p = 5.08726615734E-5), but was up-regulated by pre-miR-382 (p = 1.69645597989E-5). Values are mean ± SEM from 5 independent experiments (n = 5), compared with that in control (oligo control).
- At protein level, pre-miR-382 decreased the expression of DRD1 (p = 2.44969469509E-5) and DeltaFosB (p = 0.0008). In contrast, the expression of DRD1 (p = 0.00089) and DeltaFosB (p = 0.00149) was increased by LNA-anti-miR-382 (Anti-miR-382). Values are mean ± SEM from 5 independent experiments (n = 5), compared with that in oligo control.
- Representative Western blots of DRD1 and DeltaFosB.
- pre-miR-382 decreased, whereas Anti-miR-382 increased the expression of Drd1 (p = 0.00416 and 0.00039) and DeltaFosB (p = 7.21292762637E-5 and 0.00027) at mRNA level. Values are mean ± SEM from 5 independent experiments (n = 5), compared with that in oligo control.
- DRD1 protein was knocked-down by its siRNAs (DRD1-siRNA) (p = 0.00137). Values are mean ± SEM from 3 independent experiments (n = 3), compared with that in siRNA control.
- DeltaFosB protein was decreased via knocking-down of DRD1 by its siRNAs (p = 0.00817). Values are mean ± SEM from 3 independent experiments (n = 3), compared with that in siRNA control.
