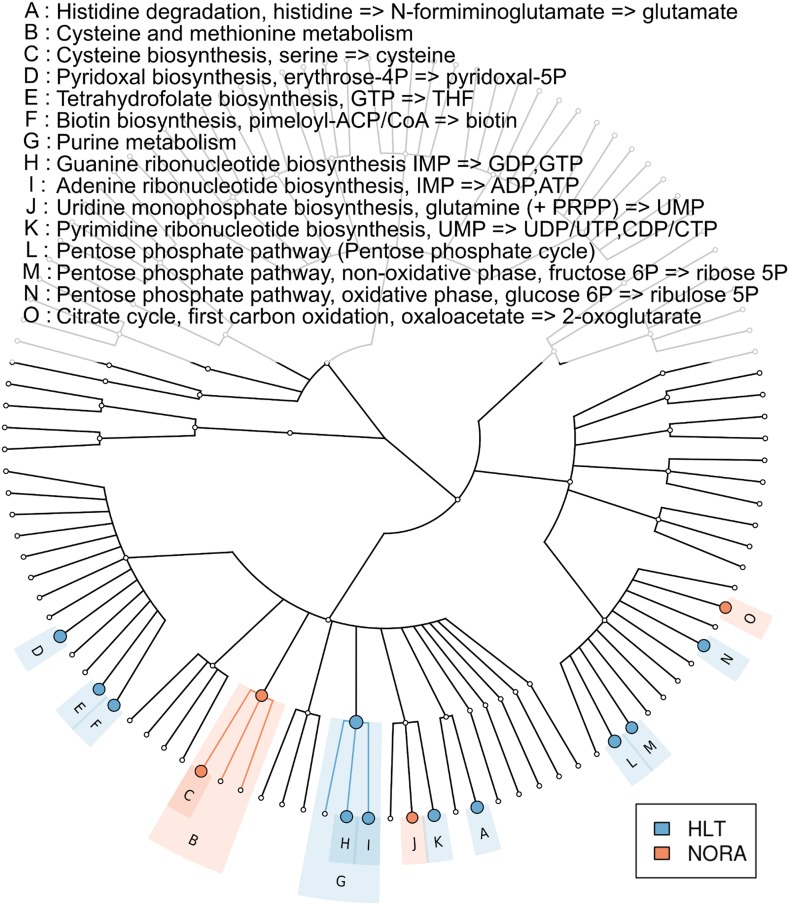
Figure 4. Metabolic pathway representation in the microbiome of healthy and NORA subjects.
HUMAnN (Abubucker et al., 2012) was applied to metagenomic reads (paired-end, 100 nt, Illumina platform) from NORA subjects (n = 14) and healthy controls (n = 5) to quantitate the abundances of hierarchically related KEGG modules in these samples (‘Materials and methods’ and Supplementary file 1A). LEfSe (Segata et al., 2011) was used to find statistically significant differences between groups at an alpha cutoff of 0.001 and an effect size cutoff of 2.0. Results shown here are highly significant (p<0.001) and represent large differences between groups. Modules highlighted in red are over-abundant in NORA samples while modules highlighted in blue are over-abundant in healthy samples. Prevotella-dominated NORA metagenomes have a dearth of genes encoding vitamin and purine metabolizing enzymes, and an excess of cysteine metabolizing enzymes.
DOI: http://dx.doi.org/10.7554/eLife.01202.019