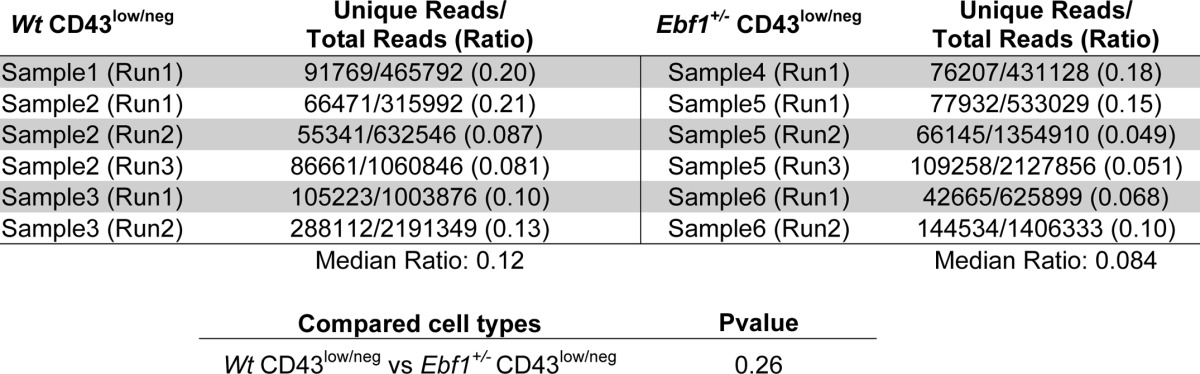
TABLE 3.
Heterozygote deletion of Ebf1 does not result in significantly decreased complexity in VDJ rearrangements in pre-B-cells
The table displays data from sequencing analysis of VDJ region complexity in the bone marrow compartment of pre-B-cells (Lin−B220−CD19+CD43low/negIgM−) from Wt and Ebf1+/− BM. The data indicate the number of reads containing a genomic J segment sequence and the number of unique sequences as judged by sequence analysis of 30 bp downstream of the genomic J segment. The data were collected from four mice (2 Wt and 2 Ebf1+/−), where the cells from each femur were sorted separately from each other. Cells from two bones from one Wt and one Ebf1+/− mouse and one bone from the second mouse of each genotype were used for the generation of three libraries from each phenotype. The libraries were analyzed in the indicated number of sequence runs. Statistical analysis was performed using the Mann-Whitney U test.