Figure 3.
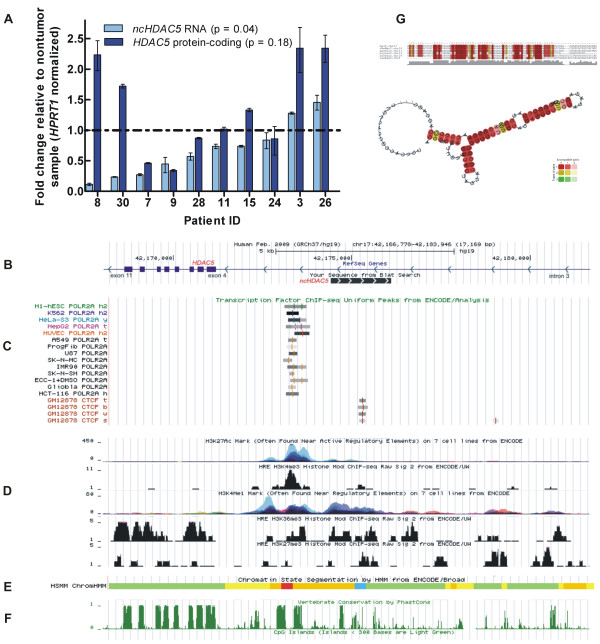
Characterization of the intronic lncRNA expressed from the HDAC5 locus. (A) Relative abundances of the ncHDAC5 lncRNA (light blue) and of the HDAC5 protein-coding mRNA (dark blue) are shown as fold change in the tumor relative to the matched nontumor sample for each of ten ccRCC patients. Patients are order according to the fold change of the ncHDAC5. (B-F) Regulatory and conserved elements from the ENCODE database are shown at the genomic region of the HDAC5 protein-coding gene from intron 3 to intron 11. Arrowheads in (B) show the opposing directions of transcription of the HDAC5 and the ncHDAC5 RNAs. In (C) the RNA Polymerase II binding sites measured in 14 cell lines, and the CTCF transcriptional repressor insulator binding site are shown. In (D) the histone modification marks H3K27ac, H3K4me3, H3K4me1, H3K36me3 and H3K27me3 are shown. In (E) the HMM histone state segmentation annotation is shown, comprising a predicted active promoter (red), a strong enhancer (orange) and an insulator (blue) region. In (F) the vertebrate conservation and the CpG islands tracks are shown (no marks detected in the latter). (G) The most stable conserved secondary structure predicted by the RNAz tool (P = 0.99) for a segment within ncHDAC5. The segment spans 110 nt along the 1.7 kb-long lncRNA transcribed in the antisense direction in the HDAC5 locus.