Figure 2.
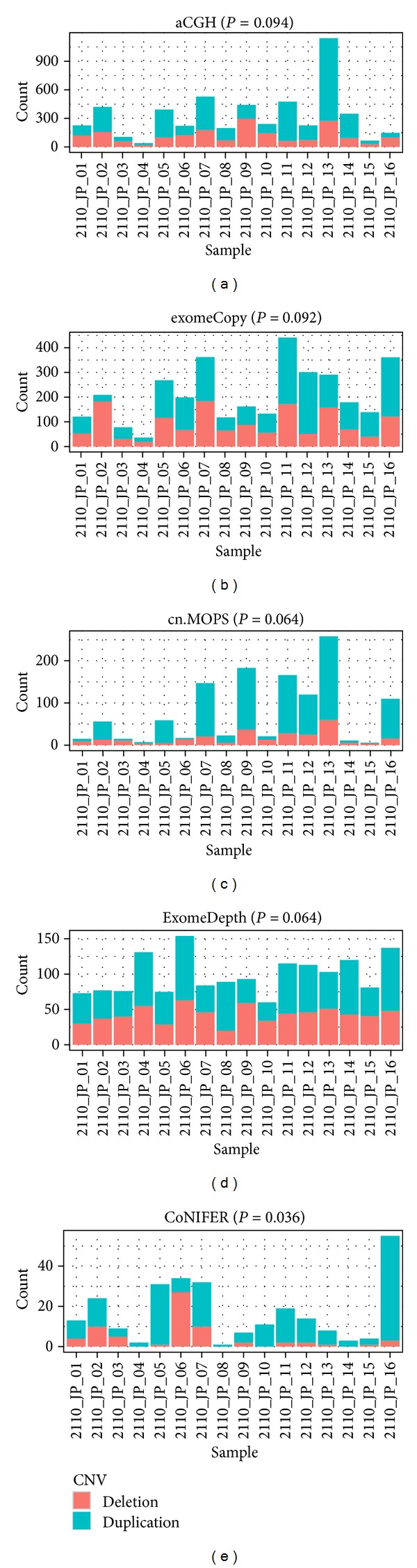
Barplot of duplication and deletion CNVs detected from each sample by five methods. The P value beside each method name was calculated by paired Wilcoxon signed rank tests following FDR correction. It indicated the detection bias between duplication and deletion CNVs of that method. Array CGH and exomeCopy showed unbiased duplication and deletion while CoNIFER had strong bias toward duplication. Cn.MOPS and ExomeDepth showed marginal bias toward duplication.
