Figure 1.
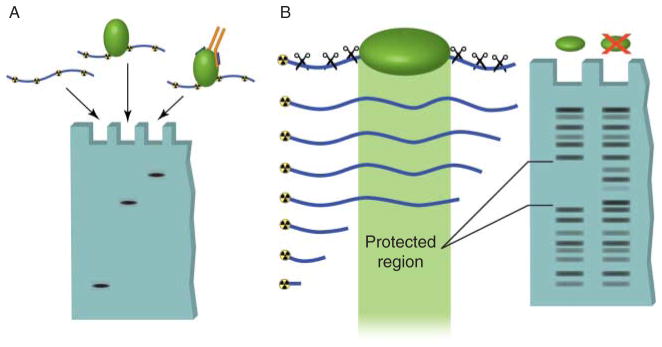
In vitro techniques used to detect DNA–protein interactions. (A) Electromobility shift assays can be used to determine direct binding between a specific sequence of radioactively labeled DNA and a purified protein. Unbound DNA, termed free probe, migrates at a relatively low molecular weight in the agarose gel. Binding of protein to this sequence results in the DNA band shifting to a high molecular weight region. Addition of an antibody that recognizes the bound protein causes an even greater shift in mobility, called supershifting. This assay can also be used with protein complexes to detect indirect protein–DNA interactions. (B) DNase footprinting assays allow identification of regions of DNA bound by proteins. A DNA oligomer is radioactively labeled on one end and mixed with the protein of interest. The DNA is then digested by a DNA endonuclease (DNase). The regions of DNA that are bound by proteins are protected from digestion. When the DNA is run out on a gel, the protected region shows up as a break in the laddering produced by DNase digestion.
