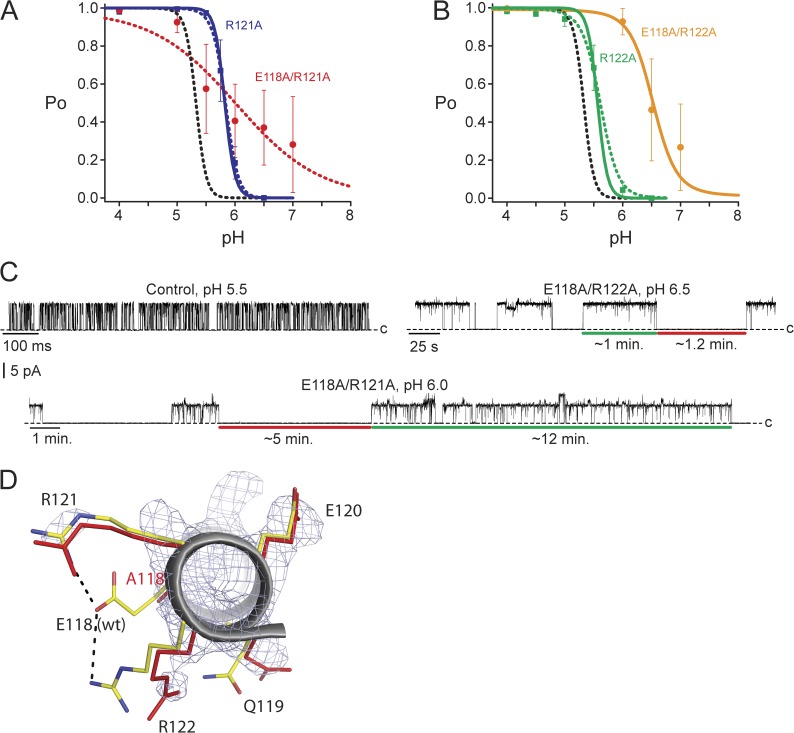
Figure 2.
Arginine residues may contribute to the intrinsic gating equilibrium as well as to the pKa values of nearby proton-binding sites. (A) R121A Po versus pH relationship (blue squares) was modeled either by changing relative to control only the intrinsic gating (solid blue line; model fit 1 in Table 1) or by also changing pKa values (dashed blue line; model fit 2 in Table 1). The E118A/R121A pH dependence (red circles) was not accurately established, and the red dotted line has no theoretical meaning. (B) R122A Po versus pH relationship (green squares) was modeled either by changing relative to control only the intrinsic gating (solid green line; model fit 1 in Table 1) or by also changing pKa values (dashed green line; model fit 2 in Table 1). The E118A/R122A pH dependence (orange circles) was not accurately established, and the orange line has no theoretical meaning. Data in A and B are the mean ± SEM from three to eight bilayers, and the model for control is shown as a black dashed line. MWC model fit parameters used in A and B are given in Table 1. (C) Example recordings of E71A control, E118A/R122A, and E118A/R121A at 100 mV and filtered offline to 50 Hz (or 20 Hz for E118A/R121A) for display. (D) The intracellular end of the TM2 helix (gray ribbon) of the E118A KcsA mutant aligned with the same region from KcsA wild type (Protein Data Bank accession no. 1K4C). The residues of interest are displayed as red sticks (E118A mutant) or with carbons in yellow, nitrogens in blue, and oxygens in red (wild type). The electron density map for the E118A KcsA mutant is shown contoured at 1.2 σ (Table 2). R121 was more disordered compared with wild type, and R122 assumed a new conformation.