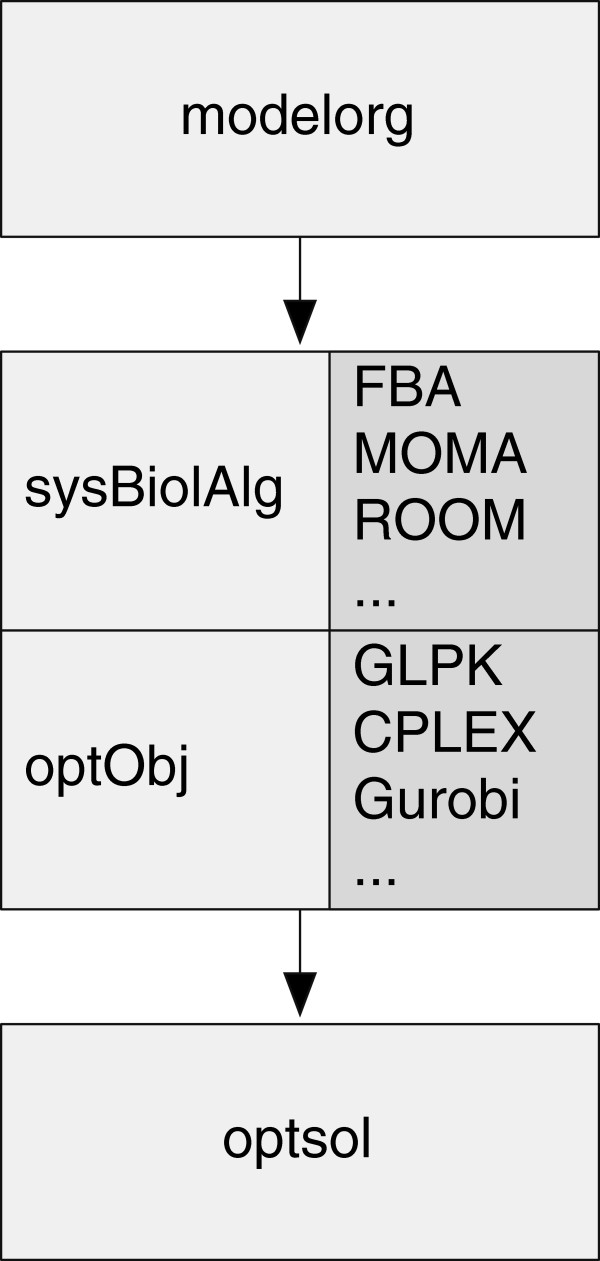
Figure 1.
Main classes in sybil. Class modelorg serves as sybil’s representation of a metabolic model. Instances of that class harbour the stoichiometric matrix, reaction names and properties, metabolites names, and gene to reaction associations. An instance of class optObj is basically a pointer to the problem object generated by the mathematical programming software. Class optObj is used for communication to the solver software. It provides methods to create, modify and solve optimisation problems, independent of the used solver software. Class sysBiolAlg holds concrete instances of class optObj which are prepared for a specific model to use with a particular algorithm (e. g. FBA or MOMA). Class sysBiolAlg is the entry point for new algorithms in sybil. The default constructor generates problem objects. Classes extending class sysBiolAlg only need to describe a specific algorithm as a formal optimisation problem. Knowledge about the details of the solver software is not required. Class optsol harbors the results of an optimisation analysis and contains analysis-specific plotting methods.