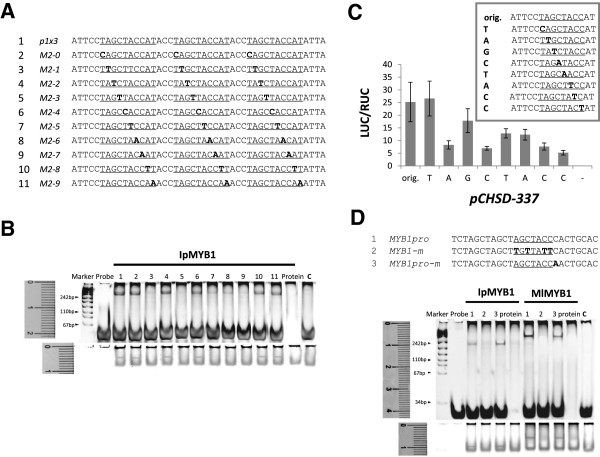
Figure 3.
Experimental tests on Motif 2. (A) Probe sequences (5 → 3’) for the EMSA tests below. The cis candidate Motif-2 (underlined) on the CHS-D us1 promoter was arranged in triplets (p1x3), and mutated sequentially in each of the probes (M2-0 to M2-9) at the sites in bold. (B) EMSA results on the binding activities to the probes of (A). The reactions were separated on a 12% SDS-PAGE gel, and shown by the DNA staining (the upper panel) and the protein staining (the lower panel) sequentially. (C) Results of dual-luciferase transient expression assays for Motif 2. The mutated promoters were labeled as T, A, G, C, T, A, C, C, whose activities were expressed as LUC/RUC. The original promoter (orig.) was included as a positive control with all three effectors presented, and the case of the promoter plus IpWDR1 and IpbHLH2 effectors (−) was shown as a control for the effect of IpMYB1. The error bars were based on four independent trials. (D) Binding tests of two MYB1 proteins on an IpMYB1 promoter (KC794943). The cis candidate (in bold) and its two mutated versions were designed on three probes (1–3), respectively. Their individual reactions to IpMYB1 and MlMYB1 were shown on the gel, with the c lane including the result of a binding test between IpbHLH2 and MYB1pro as a control for specificity. The gel setting followed those of (B).