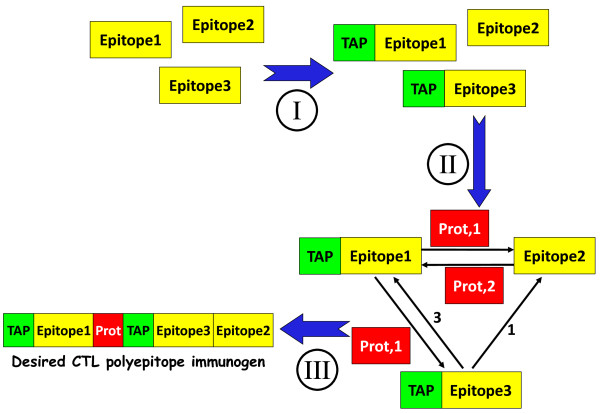
Figure 1.
PolyCTLDesigner workflow. (I) Prediction of affinity of peptides binding to TAP and addition of up to 3 N-terminal flanking amino acid residues (when necessary); (II) selection of optimal spacer sequence for each peptide pair (the optimal spacer, selected by the ranking function for each peptide pair, should meet the following criteria: it should provide formation of proteasomal cleavage site at the C-terminus of the first peptide; it should provide the least number of non-target junctional epitopes; and it should have the shortest possible length) and construction of directed weighted graph with nodes corresponding to target epitopes and with edges corresponding to allowed variants of their combinations; and (III) construction of desired polyepitope immunogen amino acid sequence which is determined as the longest simple path with a minimal weight.