Table 1. Data-collection and refinement summary.
Values in parentheses are for the highest resolution shell.
| Data-collection statistics | |
| Wavelength (Å) | 1.77 |
| Space group | P212121 |
| Unit-cell parameters (Å) | a = 93.47, b = 114.05, c = 302.52 |
| Resolution range (Å) | 49.64–2.03 (2.14–2.03) |
| Total No. of observations | 3383622 (271866) |
| No. of unique reflections | 198586 (26784) |
| Multiplicity | 17.0 (10.2) |
| Completeness (%) | 95.2 (88.9) |
| R merge † | 0.130 (0.584) |
| R meas(I)‡ | 0.142 (0.628) |
| R p.i.m.(I)‡ | 0.034 (0.171) |
| 〈I/σ(I)〉 | 25.9 (3.9) |
| Refinement statistics | |
| Resolution range (Å) | 49.64–2.03 (2.08–2.03) |
| No. of unique reflections | 198460 (13278) |
| Completeness (%) | 94.9 (86.8) |
| R cryst § (%) | 21.3 (26.6) |
| R free § (%) | 24.6 (30.5) |
| No. of protein atoms | 21390 |
| No. of heteroatoms | 224 |
| No. of water molecules | 1617 |
| Wilson B factor (Å2) | 27.8 |
| Average B factors (Å2) | |
| Overall | 31.9 |
| Protein atoms | 32.1 |
| Water molecules | 35.4 |
| Ligand (uracil) | 26.8 |
| Potassium ions | 37.1 |
| Chloride ions | 33.7 |
| Coordinate error (ESU) | 0.13 |
| Correlation coefficient, F o − F c | 0.93 |
| Correlation coefficient, F o − F c,free | 0.92 |
| Ramachandran plot, residues in (%) | |
| Allowed region | 99.7 |
| Disallowed region | 0.30 |
| MolProbity scores | |
| Clashscore | 1.23 [100th percentile] |
| Overall score | 0.98 [100th percentile] |
R
merge = 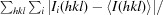
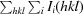 .
.
R
meas and R
p.i.m. were calculated with SCALA (Evans, 2006 ▶) in the CCP4 suite (Winn et al., 2011 ▶) using unmerged and unscaled data pre-processed by XDS (Kabsch, 2010a
▶,b
▶). R
meas is a merging R factor independent of data redundancy, while R
p.i.m. provides the precision of the averaged measurement, which improves with higher multiplicity (Weiss, 2001 ▶). R
meas = 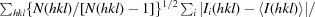
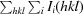 . R
p.i.m. =
. R
p.i.m. = 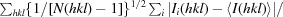
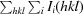
The data included in the R free set (5%) were excluded from refinement.
