FIG. 2.
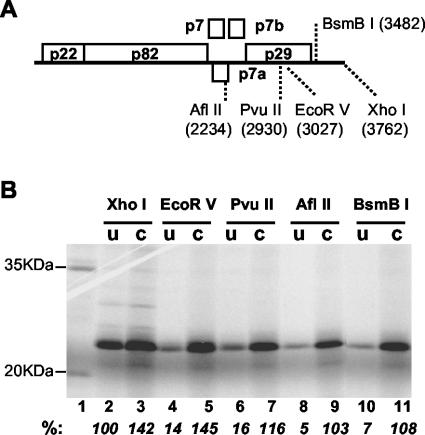
Effects of 3′ truncations on translation of TNV-D RNA in vitro. (A) Genome organization of TNV-D RNA. Restriction enzyme sites used for truncation are shown with the base number in parentheses. (B) Translation products of capped (C) or uncapped (U) TNV-D RNA truncated at the indicated restriction enzyme sites. The prominent band is p22. The predicted 104-kDa readthrough product (p22 plus p82) was not detected under these translation conditions. Lane 1 contains translation products of BMV RNAs. The migration positions (in kilodaltons) of these translation products are shown to the left of the gel. Translation was performed in wheat germ extract (Promega) with 0.2 pmol of RNA and [35S]methionine in a 25-μl reaction mixture at 25°C for 1 h. Products were separated on a sodium dodecyl sulfate-10% polyacrylamide gel and detected with a STORM 840 PhosphorImager and quantified by ImageQuant 5.2 (Amersham) software.